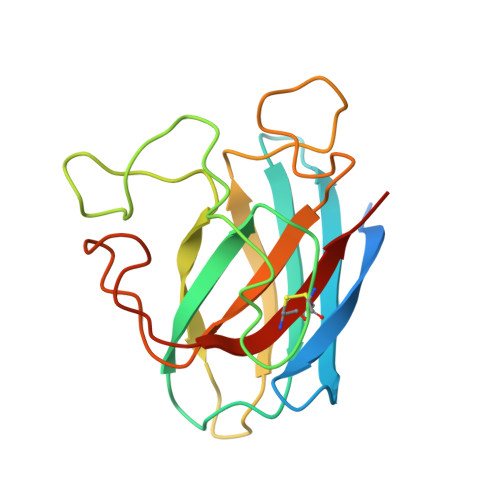
A prokaryotic superoxide dismutase paralog lacking two Cu ligands: from largely unstructured in solution to ordered in the crystal.
Banci, L., Bertini, I., Calderone, V., Cramaro, F., Del Conte, R., Fantoni, A., Mangani, S., Quattrone, A., Viezzoli, M.S.(2005) Proc Natl Acad Sci U S A 102: 7541-7546
- PubMed: 15897454
- DOI: https://doi.org/10.1073/pnas.0502450102
- Primary Citation of Related Structures:
1S4I, 1U3N - PubMed Abstract:
Little is known about prokaryotic homologs of Cu,Zn superoxide dismutase (SOD), an enzyme highly conserved among eukaryotic species. In 138 Archaea and Bacteria genomes, 57 of these putative homologs were found, 11 of which lack at least one of the metal ligands. Both the solution and the crystal structures of the SOD-like protein from Bacillus subtilis, lacking two Cu ligands and found to be enzymatically inactive, were determined. In solution, the protein is monomeric. The available nuclear Overhauser effects, together with chemical-shift index values, allowed us to define and to recognize the typical Cu,Zn SOD Greek beta-barrel but with largely unstructured loops (which, therefore, sample a wide range of conformations). On the contrary, in the crystal structure (obtained in the presence of slight excess of Zn), the protein is well structured and organized in covalent dimers held by a symmetric bridge consisting of a Zn ion bound to an Asp-His dyad in a tetrahedral geometry. Couples of dimers held by hydrophobic interactions and H bonds are further organized in long chains. The order/disorder transition is discussed in terms of metal binding and physical state.
- Department of Chemistry, University of Florence, Via Luigi Sacconi 6, 50019 Sesto, Florence, Italy.
Organizational Affiliation: