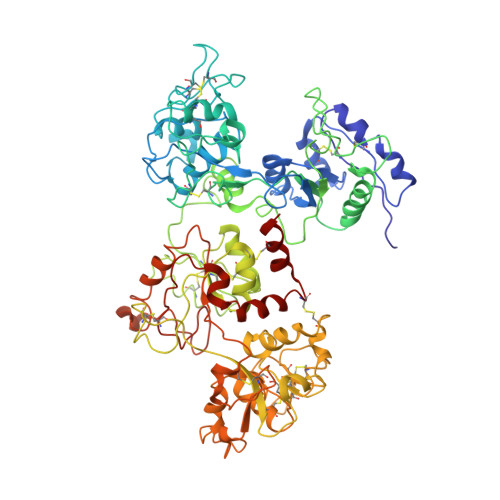
Tertiary structural changes associated with iron binding and release in hen serum transferrin: a crystallographic and spectroscopic study
Thakurta, P.G., Choudhury, D., Dasgupta, R., Dattagupta, J.K.(2004) Biochem Biophys Res Commun 316: 1124-1131
- PubMed: 15044101 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2004.02.165
- Primary Citation Related Structures:
1RYX - PubMed Abstract:
The iron binding and release of serum transferrin are pH-dependent and accompanied by a conformational change between the iron-bound (holo-) and iron-free (apo-) forms. We have determined the crystal structure of apo-hen serum transferrin (hAST) at 3.5A resolution, which is the first reported structure to date of any full molecule of an apo-serum transferrin and studied its pH-dependent iron release by UV-vis absorption and near UV-CD spectroscopy. The crystal structure of hAST shows that both the lobes adopt an open conformation and the relative orientations of the domains are different from those of apo-human serum transferrin and human apolactoferrin but similar to that of hen apo-ovotransferrin. Spectroscopic analysis reveals that in hen serum transferrin, release of the first iron starts at a pH approximately 6.5 and continues over a broad pH range (6.5-5.2). The complete release of the iron, however, occurs at pH approximately 4.0. The near UV-CD spectra show alterations in the microenvironment of the aromatic residues surrounding the iron-binding sites.
- Crystallography and Molecular Biology Division, Saha Institute of Nuclear Physics, 1/AF Bidhannagar, Kolkata 700 064, India.
Organizational Affiliation: