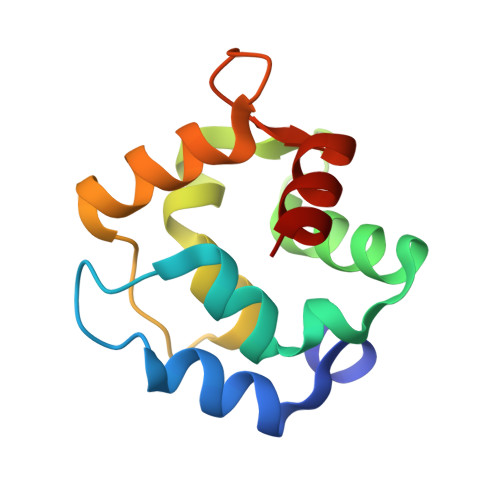
Crystal Structure of Rat Alpha-Parvalbumin at 1.05 Resolution
Bottoms, C.A., Schuermann, J.P., Agah, S., Henzl, M.T., Tanner, J.J.(2004) Protein Sci 13: 1724-1734
- PubMed: 15169955 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.03571004
- Primary Citation Related Structures:
1RWY - PubMed Abstract:
The crystal structure of rat alpha-parvalbumin has been determined at 1.05 Angstrom resolution, using synchrotron data collected at Advanced Photon Source beamline 19-ID. After refinement with SHELX, employing anisotropic displacement parameters and riding hydrogen atoms, R = 0.132 and R(free) = 0.162. The average coordinate estimated standard deviations are 0.021 Angstrom and 0.038 Angstrom for backbone atoms and side-chain atoms, respectively. Besides providing a more precise view of the alpha-isoform than previously available, these data permit comparison with the 0.91 Angstrom structure determined for pike beta-parvalbumin. Visualization of the anisotropic displacement parameters as thermal ellipsoids yields insight into the atomic motion within the Ca(2+)-binding sites. The asymmetric unit includes three parvalbumin (PV) molecules. Interestingly, the EF site in one displays uncharacteristic flexibility. The ellipsoids for Asp-92 are particularly large and non-spherical, and the shape of the Ca(2+) ellipsoid implies significant vibrational motion perpendicular to the plane defined by the four y and z ligands. The relative dearth of crystal-packing interactions in this site suggests that the heightened flexibility may be the result of diminished intermolecular contacts. The implication is that, by impeding conformational mobility, crystal-packing forces may cause serious overestimation of EF-hand rigidity. The high quality of the data permitted 11 residues to be modeled in alternative side-chain conformations, including the two core residues, Ile-97 and Leu-105. The discrete disorder observed for Ile-97 may have functional ramifications, providing a mechanism for communicating binding status between the CD and EF binding loops and between the PV metal ion-binding domain and the N-terminal AB region.
- Department of Chemistry, University of Missouri-Columbia, Columbia, MO 65211, USA.
Organizational Affiliation: