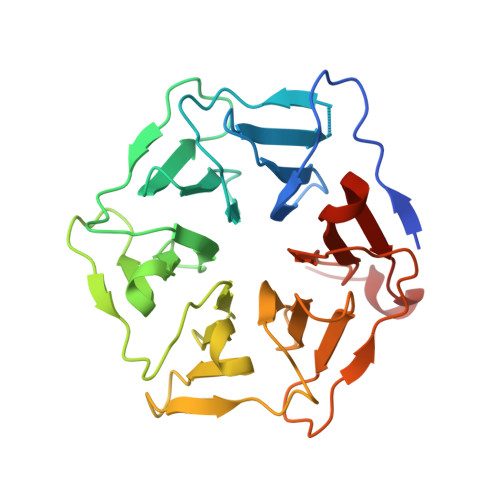
Sensor Domain of the Mycobacterium tuberculosis Receptor Ser/Thr Protein Kinase, PknD, forms a Highly Symmetric beta Propeller.
Good, M.C., Greenstein, A.E., Young, T.A., Ng, H.L., Alber, T.(2004) J Mol Biology 339: 459-469
- PubMed: 15136047 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.03.063
- Primary Citation Related Structures:
1RWI, 1RWL - PubMed Abstract:
Diverse pathogenic bacteria produce transmembrane receptor Ser/Thr protein kinases (STPKs), but little is known about the signals mediated by these "eukaryotic-like" proteins. To explore the basis for signaling in the bacterial STPK receptor family, we determined the structure of the sensor domain of Mycobacterium tuberculosis PknD. In two crystal forms, the PknD sensor domain forms a rigid, six-bladed beta-propeller with a flexible tether to the transmembrane domain. The PknD sensor domain is the most symmetric beta-propeller structure described. All residues that vary most among the blade subdomains cluster in the large "cup" motif, analogous to the ligand-binding surface in many beta-propeller proteins. These results suggest that PknD binds a multivalent ligand that signals by changing the quaternary structure of the intracellular kinase domain.
- Department of Molecular and Cell Biology, University of California, 339 Hildebrand Hall #3206, Berkeley, CA 94720-3206, USA.
Organizational Affiliation: