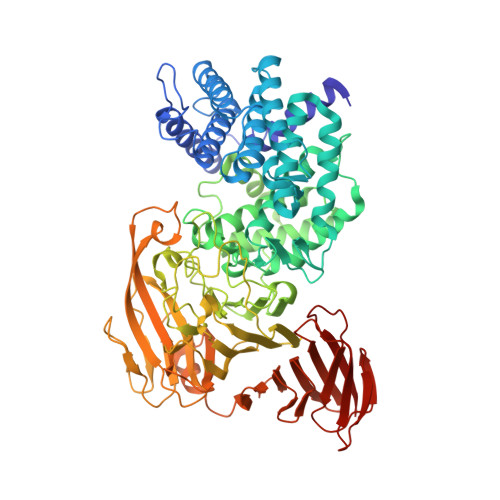
High-resolution crystal structure of Arthrobacter aurescens chondroitin AC lyase: an enzyme-substrate complex defines the catalytic mechanism
Lunin, V.V., Li, Y., Linhardt, R.J., Miyazono, H., Kyogashima, M., Kaneko, T., Bell, A.W., Cygler, M.(2004) J Mol Biology 337: 367-386
- PubMed: 15003453 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2003.12.071
- Primary Citation Related Structures:
1RW9, 1RWA, 1RWC, 1RWF, 1RWG, 1RWH - PubMed Abstract:
Chondroitin lyases (EC 4.2.2.4 and EC 4.2.2.5) are glycosaminoglycan-degrading enzymes that act as eliminases. Chondroitin lyase AC from Arthrobacter aurescens (ArthroAC) is known to act on chondroitin 4-sulfate and chondroitin 6-sulfate but not on dermatan sulfate. Like other chondroitin AC lyases, it is capable of cleaving hyaluronan. We have determined the three-dimensional crystal structure of ArthroAC in its native form as well as in complex with its substrates (chondroitin 4-sulfate tetrasaccharide, CS(tetra) and hyaluronan tetrasaccharide) at resolution varying from 1.25 A to 1.9A. The primary sequence of ArthroAC has not been previously determined but it was possible to determine the amino acid sequence of this enzyme from the high-resolution electron density maps and to confirm it by mass spectrometry. The enzyme-substrate complexes were obtained by soaking the substrate into the crystals for varying lengths of time (30 seconds to ten hours) and flash-cooling the crystals. The electron density map for crystals soaked in the substrate for as short as 30 seconds showed the substrate clearly and indicated that the ring of central glucuronic acid assumes a distorted boat conformation. This structure strongly supports the lytic mechanism where Tyr242 acts as a general base that abstracts the proton from the C5 position of glucuronic acid while Asn183 and His233 neutralize the charge on the glucuronate acidic group. Comparison of this structure with that of chondroitinase AC from Flavobacterium heparinum (FlavoAC) provides an explanation for the exolytic and endolytic mode of action of ArthroAC and FlavoAC, respectively.
- Biotechnology Research Institute, National Research Council of Canada, and Montréal Joint Centre for Structural Biology, Montréal, Québec, 6100 Royalmount Ave., Montréal, Québec, Canada H4P 2R2.
Organizational Affiliation: