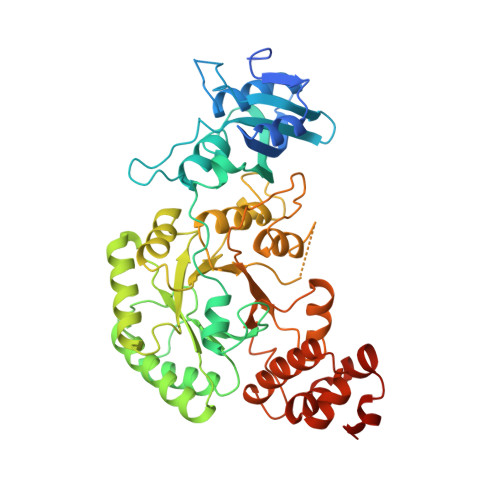
Crystal structure of the binary complex of ribulose-1,5-bisphosphate carboxylase and its product, 3-phospho-D-glycerate.
Lundqvist, T., Schneider, G.(1989) J Biological Chem 264: 3643-3646
- PubMed: 2492987 Search on PubMed
- DOI: https://doi.org/10.2210/pdb1rus/pdb
- Primary Citation Related Structures:
1RUS - PubMed Abstract:
The crystal structure of the binary complex of non-activated ribulose-1,5-bisphosphate carboxylase/oxygenase from Rhodospirillum rubrum and its product 3-phospho-D-glycerate has been determined to 2.9-A resolution. This structure determination confirms the proposed location of the active site (Schneider, G., Lindqvist, Y., Brändén, C.-I., and Lorimer, G. (1986) EMBO J. 5, 3409-3415) at the carboxyl end of the beta-strands of the alpha/beta-barrel in the carboxyl-terminal domain. One molecule of 3-phosphoglycerate is bound per active site. All oxygen atoms of 3-phosphoglycerate form hydrogen bonds to groups of the enzyme. The phosphate group interacts with the sidechains of residues Arg-288, His-321, and Ser-368, which are conserved between enzymes from different species as well as with the main chain nitrogens from residues Thr-322 and Gly-323. These amino acid residues constitute one of the two phosphate binding sites of the active site. The carboxyl group interacts with the side chains of His-287, Lys-191, and Asn-111. Implications of the activation process for the binding of 3-phosphoglycerate are discussed.
- Swedish University of Agricultural Sciences, Department of Molecular Biology, Uppsala Biomedical Center, Sweden.
Organizational Affiliation: