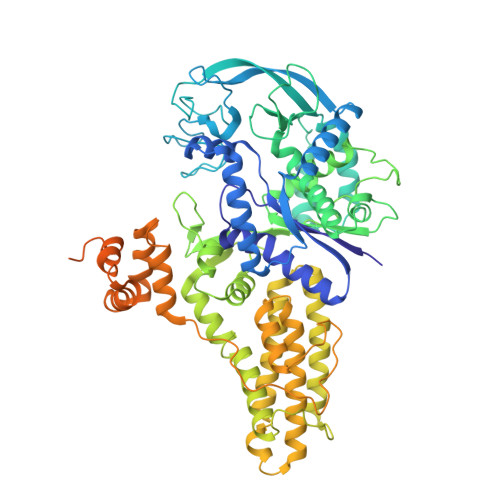
Three-dimensional structure of methionyl-tRNA synthetase from Pyrococcus abyssi.
Crepin, T., Schmitt, E., Blanquet, S., Mechulam, Y.(2004) Biochemistry 43: 2635-2644
- PubMed: 14992601
- DOI: https://doi.org/10.1021/bi0356247
- Primary Citation of Related Structures:
1RQG - PubMed Abstract:
In class 1 aminoacyl-tRNA synthetases, methionyl-tRNA synthetases (MetRS) are homodimers or monomers depending on the presence or absence of a domain appended at the C-side of the polypeptide chain. Beyond this C-domain, all MetRS display a highly conserved catalytic core with a Rossmann fold, the two halves of which are linked by a connective peptide (CP). Three-dimensional folding of CP and its putative zinc content have served as a basis to propose a division of the MetRS family into four subgroups. All subgroups but one, which is predicted to display two zincs per MetRS polypeptide, have been characterized. In the present study, the 3D structure of MetRS from Pyrococcus abyssi could be solved at 2.9 A resolution. The data obtained and atomic absorption spectroscopic measurements establish the presence of two metal ions per polypeptide chain. This finding brings strong support to the above classification. In the crystal, the C-terminal dimerization domain is disordered. This observation is thought to reflect marked flexibility of the two core moieties with respect to the C-domains in the dimer. Gel shift experiments were performed with the isolated C-terminal dimerization domain and a core monomeric MetRS, both derived from the P. abyssi enzyme. Complex formation between the C-domain and the core enzyme could not be evidenced. Moreover, association of tRNA(Met) to the core enzyme is enhanced in the presence of the C-domain. Together, these experiments suggest positive control in trans by the C-domain on recognition of tRNA by the core moiety of MetRS.
- Laboratoire de Biochimie, Unité Mixte de Recherche 7654, CNRS-Ecole Polytechnique, F-91128 Palaiseau Cedex, France.
Organizational Affiliation: