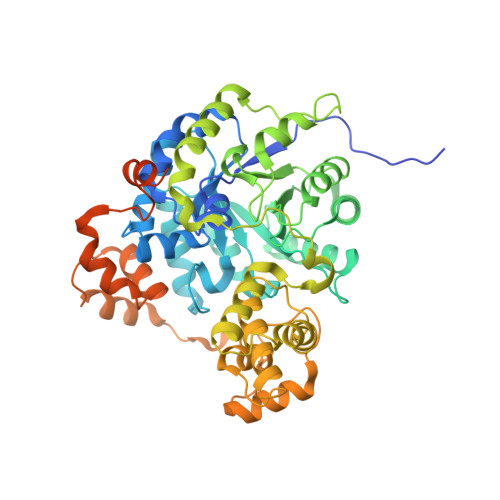
Transcarboxylase 5S structures: assembly and catalytic mechanism of a multienzyme complex subunit.
Hall, P.R., Zheng, R., Antony, L., Pusztai-Carey, M., Carey, P.R., Yee, V.C.(2004) EMBO J 23: 3621-3631
- PubMed: 15329673 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7600373
- Primary Citation Related Structures:
1RQB, 1RQE, 1RQH, 1RR2, 1S3H, 1U5J - PubMed Abstract:
Transcarboxylase is a 1.2 million Dalton (Da) multienzyme complex from Propionibacterium shermanii that couples two carboxylation reactions, transferring CO(2)(-) from methylmalonyl-CoA to pyruvate to yield propionyl-CoA and oxaloacetate. Crystal structures of the 5S metalloenzyme subunit, which catalyzes the second carboxylation reaction, have been solved in free form and bound to its substrate pyruvate, product oxaloacetate, or inhibitor 2-ketobutyrate. The structure reveals a dimer of beta(8)alpha(8) barrels with an active site cobalt ion coordinated by a carbamylated lysine, except in the oxaloacetate complex in which the product's carboxylate group serves as a ligand instead. 5S and human pyruvate carboxylase (PC), an enzyme crucial to gluconeogenesis, catalyze similar reactions. A 5S-based homology model of the PC carboxyltransferase domain indicates a conserved mechanism and explains the molecular basis of mutations in lactic acidemia. PC disease mutations reproduced in 5S result in a similar decrease in carboxyltransferase activity and crystal structures with altered active sites.
- Department of Pharmacology, Case Western Reserve University, Cleveland, OH, USA.
Organizational Affiliation: