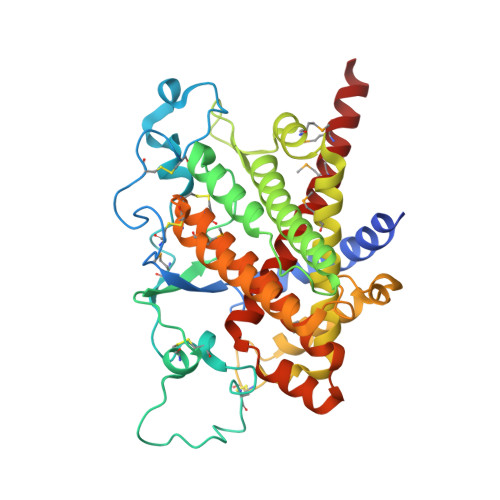
Structure of ero1p, source of disulfide bonds for oxidative protein folding in the cell.
Gross, E., Kastner, D.B., Kaiser, C.A., Fass, D.(2004) Cell 117: 601-610
- PubMed: 15163408 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(04)00418-0
- Primary Citation Related Structures:
1RP4, 1RQ1 - PubMed Abstract:
The flavoenzyme Ero1p produces disulfide bonds for oxidative protein folding in the endoplasmic reticulum. Disulfides generated de novo within Ero1p are transferred to protein disulfide isomerase and then to substrate proteins by dithiol-disulfide exchange reactions. Despite this key role of Ero1p, little is known about the mechanism by which this enzyme catalyzes thiol oxidation. Here, we present the X-ray crystallographic structure of Ero1p, which reveals the molecular details of the catalytic center, the role of a CXXCXXC motif, and the spatial relationship between functionally significant cysteines and the bound cofactor. Remarkably, the Ero1p active site closely resembles that of the versatile thiol oxidase module of Erv2p, a protein with no sequence homology to Ero1p. Furthermore, both Ero1p and Erv2p display essential dicysteine motifs on mobile polypeptide segments, suggesting that shuttling electrons to a rigid active site using a flexible strand is a fundamental feature of disulfide-generating flavoenzymes.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot 76100, Israel.
Organizational Affiliation: