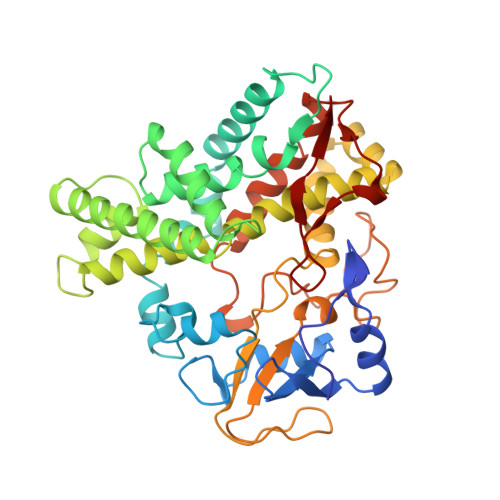
Crystal structure of nitric oxide reductase from denitrifying fungus Fusarium oxysporum.
Park, S.Y., Shimizu, H., Adachi, S., Nakagawa, A., Tanaka, I., Nakahara, K., Shoun, H., Obayashi, E., Nakamura, H., Iizuka, T., Shiro, Y.(1997) Nat Struct Biol 4: 827-832
- PubMed: 9334748 Search on PubMed
- DOI: https://doi.org/10.1038/nsb1097-827
- Primary Citation Related Structures:
1ROM, 2ROM - PubMed Abstract:
Structures of nitric oxide reductase (NOR) in the ferric resting and the ferrous CO states have been solved at 2.0 A resolution. These structures provide significant new insights into how NO is reduced in biological systems. The haem distal pocket is open to solvent, implicating this region as a possible NADH binding site. In combination with mutagenesis results, a hydrogen-bonding network from the water molecule adjacent to the iron ligand to the protein surface of the distal pocket through the hydroxyl group of Ser 286 and the carboxyl group of Asp 393 can be assigned to a pathway for proton delivery during the NO reduction reaction.
- Institute of Physical and Chemical Research (RIKEN), Saitama, Japan.
Organizational Affiliation: