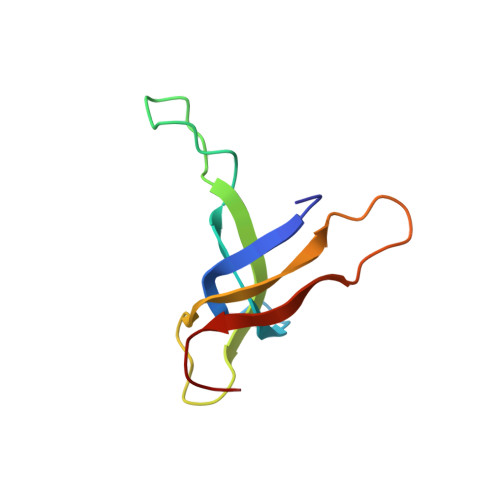
Ribosomal protein S17: characterization of the three-dimensional structure by 1H and 15N NMR.
Golden, B.L., Hoffman, D.W., Ramakrishnan, V., White, S.W.(1993) Biochemistry 32: 12812-12820
- PubMed: 8251502 Search on PubMed
- DOI: https://doi.org/10.1021/bi00210a033
- Primary Citation Related Structures:
1RIP - PubMed Abstract:
The structure of ribosomal protein S17 from Bacillus stearothermophilus was investigated by two-dimensional homonuclear and heteronuclear magnetic resonance spectroscopy. The 1H and 15N chemical shift assignments are largely complete, and a preliminary structural characterization is presented. The protein consists of five beta-strands that form a single antiparallel beta-sheet with Greek-key topology. The beta-strands are connected by several extended loops, and two of these contain residue types that are frequently seen in the RNA-binding sites of proteins. Additionally, two point mutations that affect antibiotic resistance, translational fidelity, and ribosome assembly are located in these two regions of the protein. Since these potential RNA-binding sites are distributed over a large surface of the protein, it appears that the molecule may interact with several regions of 16S rRNA.
- Department of Microbiology, Duke University Medical Center, Durham, North Carolina 27710.
Organizational Affiliation: