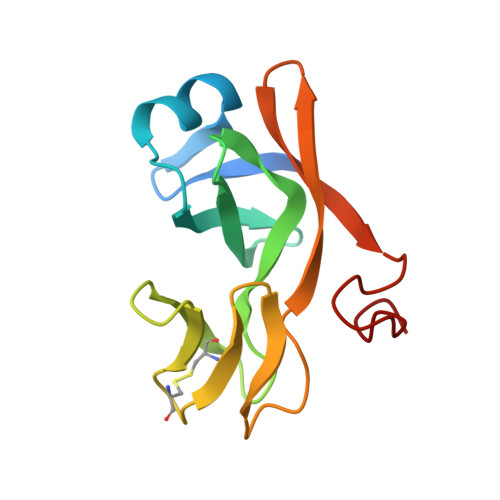
Biological identity and diversity in photosynthesis and respiration: structure of the lumen-side domain of the chloroplast Rieske protein.
Carrell, C.J., Zhang, H., Cramer, W.A., Smith, J.L.(1997) Structure 5: 1613-1625
- PubMed: 9438861 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(97)00309-2
- Primary Citation Related Structures:
1RFS - PubMed Abstract:
The cytochrome b6f complex functions in oxygenic photosynthesis as an integral membrane protein complex that mediates coupled electron transfer and proton translocation. The Rieske [2Fe-2S] protein subunit of the complex functions at the electropositive (p) membrane interface as the electron acceptor for plastoquinol and donor for the cytochrome f subunit, and may have a dynamic role in catalyzing electron and proton transfer at the membrane interface. There are significant structure/function similarities to the cytochrome bc1 complex of the respiratory chain. The 1.83 A crystal structure of a 139-residue C-terminal fragment of the Rieske [2Fe-2S] protein, derived from the cytochrome b6f complex of spinach chloroplasts, has been solved by multiwavelength anomalous diffraction. The structure of the fragment comprises two domains: a small 'cluster-binding' subdomain and a large subdomain. The [2Fe-2S] cluster-binding subdomains of the chloroplast and mitochondrial Rieske proteins are virtually identical, whereas the large subdomains are strikingly different despite a common folding topology. A structure-based sequence alignment of the b6f and bc1 groups of Rieske soluble domains is presented. The segregation of structural conservation and divergence in the cluster-binding and large subdomains of the Rieske protein correlates with the overall relatedness of the cytochrome b6f and bc1 complexes, in which redox domains in the aqueous p phase are dissimilar and those within the membrane are similar. Distinct sequences and surface charge distributions among Rieske large subdomains may provide a signature for interaction with the p-side oxidant protein and for the pH of the intraorganelle compartment.
- Department of Biological Sciences, Purdue University, West Lafayette, IN 47907-1392, USA.
Organizational Affiliation: