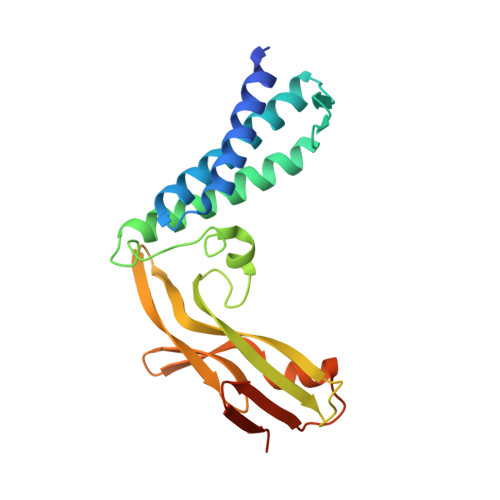
Crystal Structure of the E2 Transactivation Domain of Human Papillomavirus Type 11 Bound to a Protein Interaction Inhibitor
Wang, Y., Coulombe, R., Cameron, D.R., Thauvette, L., Massariol, M.-J., Amon, L.M., Fink, D., Titolo, S., Welchner, E., Yoakim, C., Archambault, J., White, P.W.(2004) J Biological Chem 279: 6976-6985
- PubMed: 14634007
- DOI: https://doi.org/10.1074/jbc.M311376200
- Primary Citation Related Structures:
1R6K, 1R6N - PubMed Abstract:
Interaction between the E2 protein and E1 helicase of human papillomaviruses (HPVs) is essential for the initiation of viral DNA replication. We recently described a series of small molecules that bind to the N-terminal transactivation domain (TAD) of HPV type 11 E2 and inhibits its interaction with E1 in vitro and in cellular assays. Here we report the crystal structures of both the HPV11 TAD and of a complex between this domain and an inhibitor, at 2.5- and 2.4-A resolution, respectively. The HPV11 TAD structure is very similar to that of the analogous domain of HPV16. Inhibitor binding caused no significant alteration of the protein backbone, but movements of several amino acid side chains at the binding site, in particular those of Tyr-19, His-32, Leu-94, and Glu-100, resulted in the formation of a deep hydrophobic pocket that accommodates the indandione moiety of the inhibitor. Mutational analysis provides functional evidence for specific interactions between Tyr-19 and E1 and between His-32 and the inhibitor. A second inhibitor molecule is also present at the binding pocket. Although evidence is presented that this second molecule makes only weak interactions with the protein and is likely an artifact of crystallization, its presence defines additional regions of the binding pocket that could be exploited to design more potent inhibitors.
- Department of Medicinal Chemistry, Boehringer Ingelheim Pharmaceuticals Inc, Ridgefield, CT 06877, USA.
Organizational Affiliation: