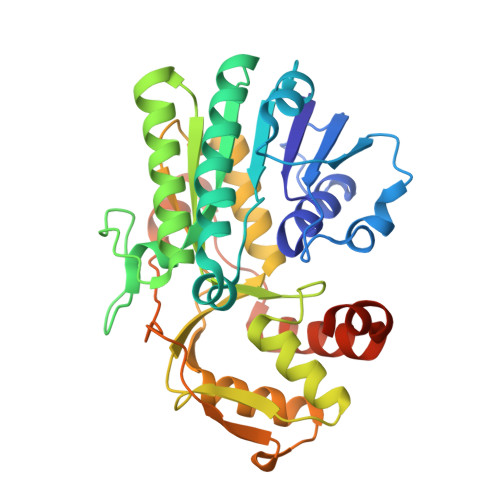
High Resolution X-ray Structure of dTDP-Glucose 4,6-Dehydratase from Streptomyces venezuelae
Allard, S.T.M., Cleland, W.W., Holden, H.M.(2004) J Biological Chem 279: 2211-2220
- PubMed: 14570895 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M310134200
- Primary Citation Related Structures:
1R66, 1R6D - PubMed Abstract:
Desosamine is a 3-(dimethylamino)-3,4,6-trideoxyhexose found in some macrolide antibiotics. In Streptomyces venezuelae, there are seven genes required for the biosynthesis of this unusual sugar. One of the genes, desIV, codes for a dTDP-glucose 4,6-dehydratase, which is referred to as DesIV. The reaction mechanisms for these types of dehydratases are quite complicated with proton abstraction from the sugar 4'-hydroxyl group and hydride transfer to NAD+, proton abstraction at C-5, and elimination of the hydroxyl group at C-6 of the sugar, and finally return of a proton to C-5 and a hydride from NADH to C-6. Here we describe the cloning, overexpression, and purification, and high resolution x-ray crystallographic analysis to 1.44 A of wild-type DesIV complexed with dTDP. Additionally, for this study, a double site-directed mutant protein (D128N/E129Q) was prepared, crystallized as a complex with NAD+ and the substrate dTDP-glucose and its structure determined to 1.35 A resolution. In DesIV, the phenolate group of Tyr(151) and O(gamma) of Thr(127) lie at 2.7 and 2.6 A, respectively from the 4'-hydroxyl group of the dTDP-glucose substrate. The side chain of Asp(128) is in the correct position to function as a general acid for proton donation to the 6'-hydroxyl group while the side chain of Glu(129) is ideally situated to serve as the general base for proton abstraction at C-5. This investigation provides further detailed information for understanding the exquisite chemistry that occurs in these remarkable enzymes.
- Department of Biochemistry, University of Wisconsin, Madison, WI 53706, USA.
Organizational Affiliation: