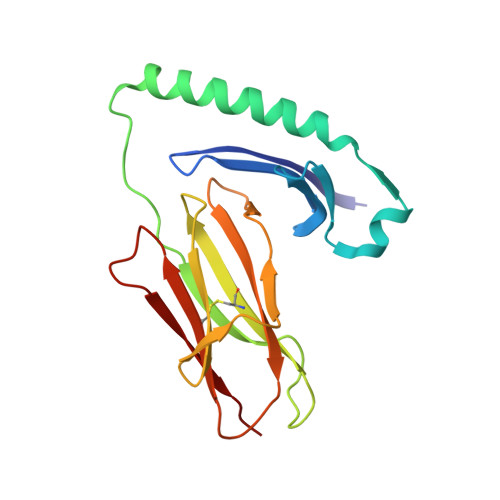
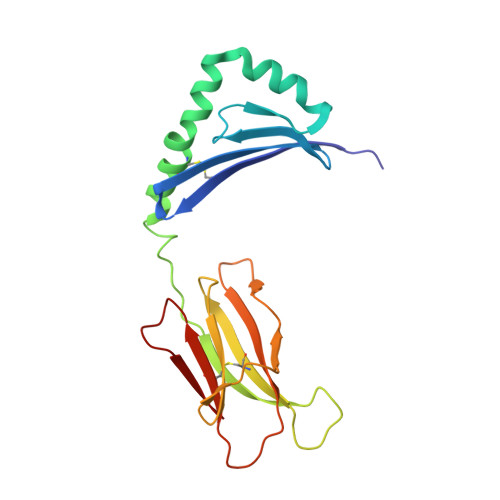
Evidence that structural rearrangements and/or flexibility during TCR binding can contribute to T cell activation.
Krogsgaard, M., Prado, N., Adams, E.J., He, X.L., Chow, D.C., Wilson, D.B., Garcia, K.C., Davis, M.M.(2003) Mol Cell 12: 1367-1378
- PubMed: 14690592
- DOI: https://doi.org/10.1016/s1097-2765(03)00474-x
- Primary Citation of Related Structures:
1R5V, 1R5W - PubMed Abstract:
While in many cases the half-life of T cell receptor (TCR) binding to a particular ligand is a good predictor of activation potential, numerous exceptions suggest that other physical parameter(s) must also play a role. Accordingly, we analyzed the thermodynamics of TCR binding to a series of peptide-MHC ligands, three of which are more stimulatory than their stability of binding would predict. Strikingly, we find that during TCR binding these outliers show anomalously large changes in heat capacity, an indicator of conformational change or flexibility in a binding interaction. By combining the values for heat capacity (DeltaCp) and the half-life of TCR binding (t(1/2)), we find that we can accurately predict the degree of T cell stimulation. Structural analysis shows significant changes in the central TCR contact residue of the peptide-MHC, indicating that structural rearrangements within the TCR-peptide-MHC interface can contribute to T cell activation.
- Howard Hughes Medical Institute, Stanford University School of Medicine, Stanford, CA 94305, USA.
Organizational Affiliation: