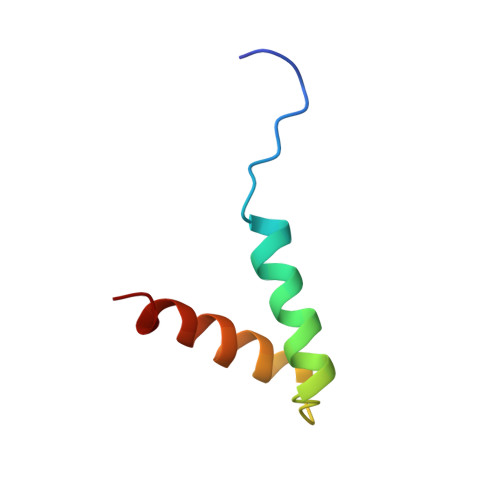
The molecular basis for protein kinase A anchoring revealed by solution NMR.
Newlon, M.G., Roy, M., Morikis, D., Hausken, Z.E., Coghlan, V., Scott, J.D., Jennings, P.A.(1999) Nat Struct Biol 6: 222-227
- PubMed: 10074940 Search on PubMed
- DOI: https://doi.org/10.1038/6663
- Primary Citation Related Structures:
1R2A - PubMed Abstract:
Compartmentalization of signal transduction enzymes into signaling complexes is an important mechanism to ensure the specificity of intracellular events. Formation of these complexes is mediated by specialized protein motifs that participate in protein-protein interactions. The adenosine 3',5'-cyclic monophosphate (cAMP)-dependent protein kinase (PKA) is localized through interaction of the regulatory (R) subunit dimer with A-kinase-anchoring proteins (AKAPs). We now report the solution structure of the type II PKA R-subunit fragment RIIalpha(1-44), which encompasses both the AKAP-binding and dimerization interfaces. This structure incorporates an X-type four-helix bundle dimerization motif with an extended hydrophobic face that is necessary for high-affinity AKAP binding. NMR data on the complex between RIIalpha(1-44) and an AKAP fragment reveals extensive contacts between the two proteins. Interestingly, this same dimerization motif is present in other signaling molecules, the S100 family. Therefore, the X-type four-helix bundle may represent a conserved fold for protein-protein interactions in signal transduction.
- Department of Chemistry and Biochemistry, University of California, San Diego, La Jolla 92093-0359, USA.
Organizational Affiliation: