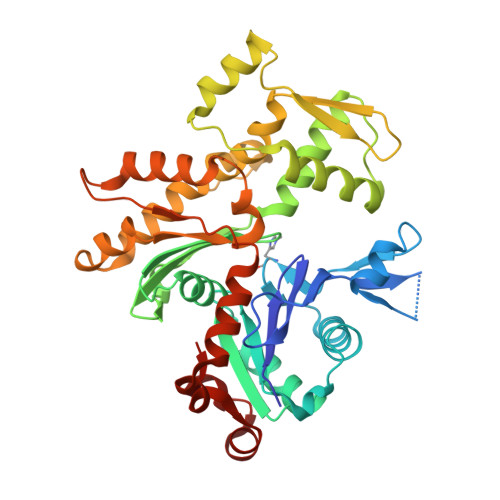
Trisoxazole macrolide toxins mimic the binding of actin-capping proteins to actin
Klenchin, V.A., Allingham, J.S., King, R., Tanaka, J., Marriott, G., Rayment, I.(2003) Nat Struct Biol 10: 1058-1063
- PubMed: 14578936 Search on PubMed
- DOI: https://doi.org/10.1038/nsb1006
- Primary Citation Related Structures:
1QZ5, 1QZ6 - PubMed Abstract:
Marine macrolide toxins of trisoxazole family target actin with high affinity and specificity and have promising pharmacological properties. We present X-ray structures of actin in complex with two members of this family, kabiramide C and jaspisamide A, at a resolution of 1.45 and 1.6 A, respectively. The structures reveal the absolute stereochemistry of these toxins and demonstrate that their trisoxazole ring interacts with actin subdomain 1 while the aliphatic side chain is inserted into the hydrophobic cavity between actin subdomains 1 and 3. The binding site is essentially the same as the one occupied by the actin-capping domain of the gelsolin superfamily of proteins. The structural evidence suggests that actin filament severing and capping by these toxins is also analogous to that of gelsolin. Consequently, these macrolides may be viewed as small molecule biomimetics of an entire class of actin-binding proteins.
- Department of Biochemistry, University of Wisconsin at Madison, 433 Babcock Drive, Madison, Wisconsin 53706, USA.
Organizational Affiliation: