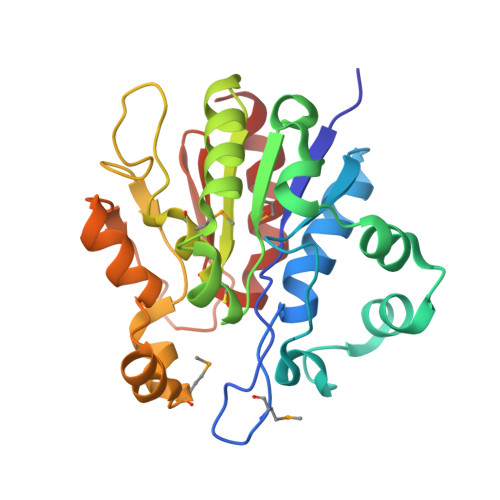
Crystal structure of the YDR533c S. cerevisiae protein, a class II member of the Hsp31 family
Graille, M., Quevillon-Cheruel, S., Leulliot, N., Zhou, C.Z., de la Sierra Gallay, I.L., Jacquamet, L., Ferrer, J.L., Liger, D., Poupon, A., Janin, J., van Tilbeurgh, H.(2004) Structure 12: 839-847
- PubMed: 15130476 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.02.030
- Primary Citation Related Structures:
1QVV, 1QVW, 1QVZ - PubMed Abstract:
The ORF YDR533c from Saccharomyces cerevisiae codes for a 25.5 kDa protein of unknown biochemical function. Transcriptome analysis of yeast has shown that this gene is activated in response to various stress conditions together with proteins belonging to the heat shock family. In order to clarify its biochemical function, we determined the crystal structure of YDR533c to 1.85 A resolution by the single anomalous diffraction method. The protein possesses an alpha/beta hydrolase fold and a putative Cys-His-Glu catalytic triad common to a large enzyme family containing proteases, amidotransferases, lipases, and esterases. The protein has strong structural resemblance with the E. coli Hsp31 protein and the intracellular protease I from Pyrococcus horikoshii, which are considered class I and class III members of the Hsp31 family, respectively. Detailed structural analysis strongly suggests that the YDR533c protein crystal structure is the first one of a class II member of the Hsp31 family.
- Laboratoire d'Enzymologie et Biochimie Structurales, CNRS-UPR 9063, Bâtiment 34, 1 Avenue de la Terrasse, 91198 Gif sur Yvette, France.
Organizational Affiliation: