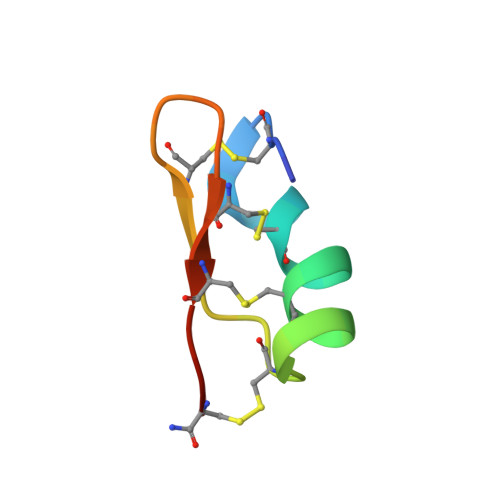
Structural and functional consequences of the presence of a fourth disulfide bridge in the scorpion short toxins: solution structure of the potassium channel inhibitor HsTX1.
Savarin, P., Romi-Lebrun, R., Zinn-Justin, S., Lebrun, B., Nakajima, T., Gilquin, B., Menez, A.(1999) Protein Sci 8: 2672-2685
- PubMed: 10631983 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.8.12.2672
- Primary Citation Related Structures:
1QUZ - PubMed Abstract:
We have determined the three-dimensional structure of the potassium channel inhibitor HsTX1, using nuclear magnetic resonance and molecular modeling. This protein belongs to the scorpion short toxin family, which essentially contains potassium channel blockers of 29 to 39 amino acids and three disulfide bridges. It is highly active on voltage-gated Kv1.3 potassium channels. Furthermore, it has the particularity to possess a fourth disulfide bridge. We show that HsTX1 has a fold similar to that of the three-disulfide-bridged toxins and conserves the hydrophobic core found in the scorpion short toxins. Thus, the fourth bridge has no influence on the global conformation of HsTX1. Most residues spatially analogous to those interacting with voltage-gated potassium channels in the three-disulfide-bridged toxins are conserved in HsTX1. Thus, we propose that Tyr21, Lys23, Met25, and Asn26 are involved in the biological activity of HsTX1. As an additional positively charged residue is always spatially close to the aromatic residue in toxins blocking the voltage-gated potassium channels, and as previous mutagenesis experiments have shown the critical role played by the C-terminus in HsTX1, we suggest that Arg33 is also important for the activity of the four disulfide-bridged toxin. Docking calculations confirm that, if Lys23 and Met25 interact with the GYGDMH motif of Kv1.3, Arg33 can contact Asp386 and, thus, play the role of the additional positively charged residue of the toxin functional site. This original configuration of the binding site of HsTX1 for Kv1.3, if confirmed experimentally, offers new structural possibilities for the construction of a molecule blocking the voltage-gated potassium channels.
- CEA, Département d'Ingénierie et d'Etude des Protéines, Gif-sur-Yvette, France.
Organizational Affiliation: