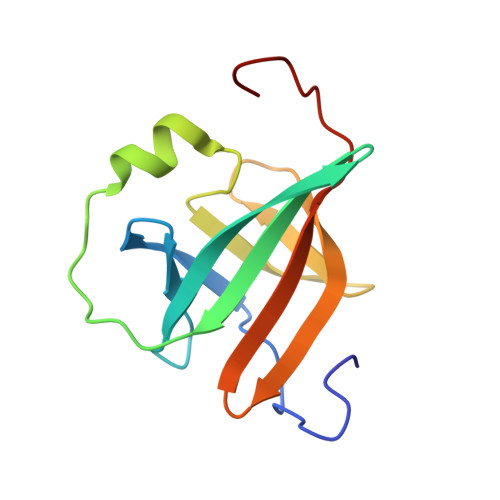
Backbone dynamics and solution structure refinement of the 15N-labeled human oncogenic protein p13MTCP1: comparison with X-ray data.
Guignard, L., Padilla, A., Mispelter, J., Yang, Y.S., Stern, M.H., Lhoste, J.M., Roumestand, C.(2000) J Biomol NMR 17: 215-230
- PubMed: 10959629
- DOI: https://doi.org/10.1023/a:1008386110930
- Primary Citation Related Structures:
1QTT, 1QTU - PubMed Abstract:
Two related oncogenes, TCL1 and MTCP1, are overexpressed in certain T-cell prolymphocytic leukemias as a result of chromosomal rearrangements that involve the translocation of one T-cell receptor gene to either chromosome 14q32 or Xq28, respectively. The human oncoprotein p13MTCP1 is coded by the MTCP1 gene and its primary sequence is highly and only homologous to that of p14TCL1, the product of TCL1. These two proteins likely represent the first members of a new family of oncogenic proteins. A previous model of the three-dimensional solution structure of p13MTCP1 was determined recently using exclusively homonuclear proton two-dimensional NMR methods and, almost simultaneously, high-resolution crystal structures of p13MTCP1 and p14TCL1 appeared in the literature. In order to gain more insight into the details of the solution structure, we uniformly labeled p13MTCP1 with nitrogen-15. The refined structure benefits from 520 additional NOEs, extracted from either 15N-edited 3D experiments or homonuclear 2D NOESY recorded at 800 MHz, and from a nearly complete set of phi angular restraints. Measurements of 15N spin relaxation times and heteronuclear 15N[1H]NOEs at two magnetic field strengths provided additional insights into the dynamics of the protein backbone. On the basis of these new results, a putative binding surface for this particular class of oncogenes is discussed.
- Centre de Biochimie Structurale, CNRS-UMR 9955, INSERM-U414, Université de Montpellier I, Faculté de Pharmacie, France.
Organizational Affiliation: