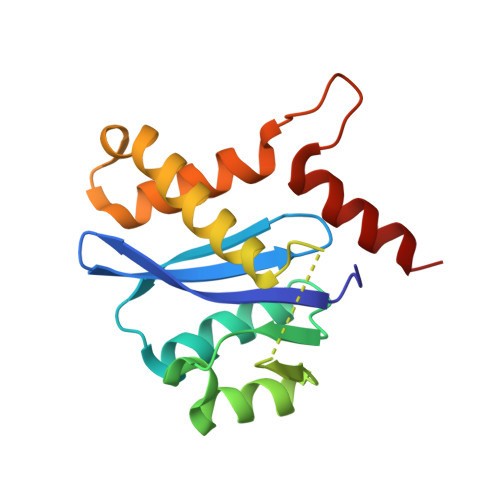
Structure of the HIV-1 integrase catalytic domain complexed with an inhibitor: a platform for antiviral drug design.
Goldgur, Y., Craigie, R., Cohen, G.H., Fujiwara, T., Yoshinaga, T., Fujishita, T., Sugimoto, H., Endo, T., Murai, H., Davies, D.R.(1999) Proc Natl Acad Sci U S A 96: 13040-13043
- PubMed: 10557269 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.96.23.13040
- Primary Citation Related Structures:
1QS4 - PubMed Abstract:
HIV integrase, the enzyme that inserts the viral DNA into the host chromosome, has no mammalian counterpart, making it an attractive target for antiviral drug design. As one of the three enzymes produced by HIV, it can be expected that inhibitors of this enzyme will complement the therapeutic use of HIV protease and reverse transcriptase inhibitors. We have determined the structure of a complex of the HIV-1 integrase core domain with a novel inhibitor, 5ClTEP, 1-(5-chloroindol-3-yl)-3-hydroxy-3-(2H-tetrazol-5-yl)-pro penone, to 2.1-A resolution. The inhibitor binds centrally in the active site of the integrase and makes a number of close contacts with the protein. Only minor changes in the protein accompany inhibitor binding. This inhibitor complex will provide a platform for structure-based design of an additional class of inhibitors for antiviral therapy.
- Laboratory of Molecular Biology, National Institute of Diabetes, Digestive and Kidney Diseases, National Institutes of Health, Bethesda, MD 20892-0560, USA.
Organizational Affiliation: