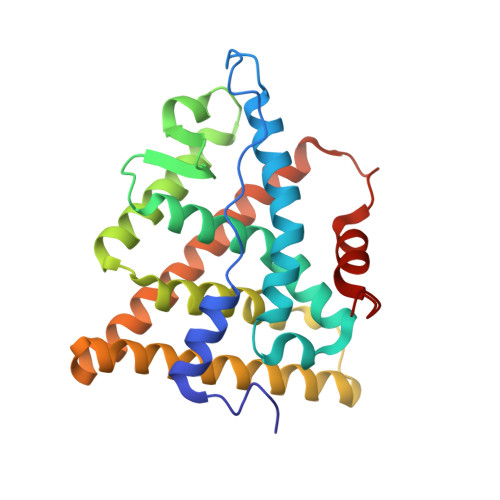
Crystal Structure of a Mutant Heralpha Ligand- Binding Domain Reveals Key Structural Features for the Mechanism of Partial Agonism
Gangloff, M., Ruff, M., Eiler, S., Duclaud, S., Wurtz, J.M., Moras, D.(2001) J Biological Chem 276: 15059
- PubMed: 11278577 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M009870200
- Primary Citation Related Structures:
1QKT, 1QKU - PubMed Abstract:
The crystal structure of a triple cysteine to serine mutant ERalpha ligand-binding domain (LBD), complexed with estradiol, shows that despite the presence of a tightly bound agonist ligand, the protein exhibits an antagonist-like conformation, similar to that observed in raloxifen and 4-hydroxytamoxifen-bound structures. This mutated receptor binds estradiol with wild type affinity and displays transcriptional activity upon estradiol stimulation, but with limited potency (about 50%). This partial activity is efficiently repressed in antagonist competition assays. The comparison with available LBD structures reveals key features governing the positioning of helix H12 and highlights the importance of cysteine residues in promoting an active conformation. Furthermore the present study reveals a hydrogen bond network connecting ligand binding to protein trans conformation. These observations support a dynamic view of H12 positioning, where the control of the equilibrium between two stable locations determines the partial agonist character of a given ligand.
- Institut de Genetique et de Biologie Moleculaire et Cellulaire, Laboratoire de Biologie et de Génomique Structurales 1, rue Laurent Fries, BP 163 67404 Illkirch, France.
Organizational Affiliation: