A New Potent Calmodulin Antagonist with Arylalkylamine Structure: Crystallographic, Spectroscopic and Functional Studies
Harmat, V., Bocskei, Z.S., Naray-Szabo, G., Bata, I., Csutor, A.S., Hermecz, I., Aranyi, P., Szabo, B., Liliom, K., Vertessy, B.G., Ovadi, J.(2000) J Mol Biology 297: 747
- PubMed: 10731425 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.3607
- Primary Citation Related Structures:
1QIV, 1QIW - PubMed Abstract:
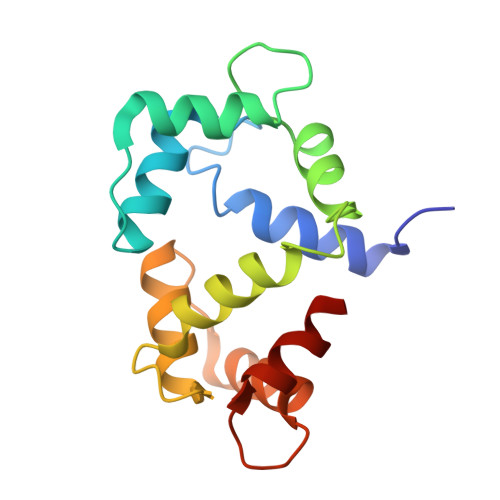
An arylalkylamine-type calmodulin antagonist, N-(3, 3-diphenylpropyl)-N'-[1-R-(3, 4-bis-butoxyphenyl)ethyl]-propylene-diamine (AAA) is presented and its complexes with calmodulin are characterized in solution and in the crystal. Near-UV circular dichroism spectra show that AAA binds to calmodulin with 2:1 stoichiometry in a Ca(2+)-dependent manner. The crystal structure with 2:1 stoichiometry is determined to 2.64 A resolution. The binding of AAA causes domain closure of calmodulin similar to that obtained with trifluoperazine. Solution and crystal data indicate that each of the two AAA molecules anchors in the hydrophobic pockets of calmodulin, overlapping with two trifluoperazine sites, i.e. at a hydrophobic pocket and an interdomain site. The two AAA molecules also interact with each other by hydrophobic forces. A competition enzymatic assay has revealed that AAA inhibits calmodulin-activated phosphodiesterase activity at two orders of magnitude lower concentration than trifluoperazine. The apparent dissociation constant of AAA to calmodulin is 18 nM, which is commensurable with that of target peptides. On the basis of the crystal structure, we propose that the high-affinity binding is mainly due to a favorable entropy term, as the AAA molecule makes multiple contacts in its complex with calmodulin.
- Department of Theoretical Chemistry, Loránd Eötvös University, Budapest 112, H-1518, Hungary. harmatv@ludens.elte.hu
Organizational Affiliation: