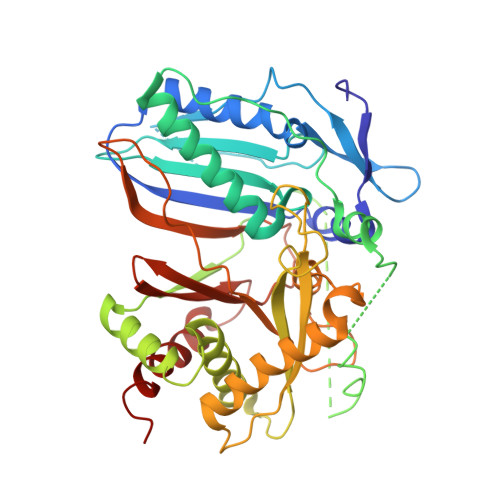
Crystal Structure of PapA5, a Phthiocerol Dimycocerosyl Transferase from Mycobacterium tuberculosis
Buglino, J., Onwueme, K.C., Ferreras, J.A., Quadri, L.E., Lima, C.D.(2004) J Biological Chem 279: 30634-30642
- PubMed: 15123643 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M404011200
- Primary Citation Related Structures:
1Q9J - PubMed Abstract:
Polyketide-associated protein A5 (PapA5) is an acyltransferase that is involved in production of phthiocerol and phthiodiolone dimycocerosate esters, a class of virulence-enhancing lipids produced by Mycobacterium tuberculosis. Structural analysis of PapA5 at 2.75-A resolution reveals a two-domain structure that shares unexpected similarity to structures of chloramphenicol acetyltransferase, dihydrolipoyl transacetylase, carnitine acetyltransferase, and VibH, a non-ribosomal peptide synthesis condensation enzyme. The PapA5 active site includes conserved histidine and aspartic acid residues that are critical to PapA5 acyltransferase activity. PapA5 catalyzes acyl transfer reactions on model substrates that contain long aliphatic carbon chains, and two hydrophobic channels were observed linking the PapA5 surface to the active site with properties consistent with these biochemical activities and substrate preferences. An additional alpha helix not observed in other acyltransferase structures blocks the putative entrance into the PapA5 active site, indicating that conformational changes may be associated with PapA5 activity. PapA5 represents the first structure solved for a protein involved in polyketide synthesis in Mycobacteria.
- Structural Biology Program, Sloan-Kettering Institute, New York, New York 10021, USA.
Organizational Affiliation: