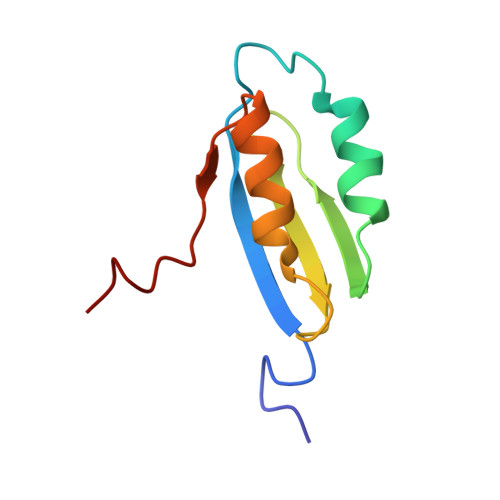
Structure and metal binding studies of the second copper binding domain of the Menkes ATPase.
Jones, C.E., Daly, N.L., Cobine, P.A., Craik, D.J., Dameron, C.T.(2003) J Struct Biol 143: 209-218
- PubMed: 14572476 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2003.08.008
- Primary Citation Related Structures:
1Q8L - PubMed Abstract:
Biological utilisation of copper requires that the metal, in its ionic forms, be meticulously transported, inserted into enzymes and regulatory proteins, and excess be excreted. To understand the trafficking process, it is crucial that the structures of the proteins involved in the varied processes be resolved. To investigate copper binding to a family of structurally related copper-binding proteins, we have characterised the second Menkes N-terminal domain (MNKr2). The structure, determined using 1H and 15N heteronuclear NMR, of the reduced form of MNKr2 has revealed two alpha-helices lying over a single beta-sheet and shows that the binding site, a Cys(X)2Cys pair, is located on an exposed loop. 1H-15N HSQC experiments demonstrate that binding of Cu(I) causes changes that are localised to conserved residues adjacent to the metal binding site. Residues in this area are important to the delivery of copper by the structurally related Cu(I) chaperones. Complementary site-directed mutagenesis of the adjacent residues has been used to probe the structural roles of conserved residues.
- The National Research Centre for Environmental Toxicology, The University of Queensland, 39 Kessels Road, Coopers Plains, Qld 4108, Australia.
Organizational Affiliation: