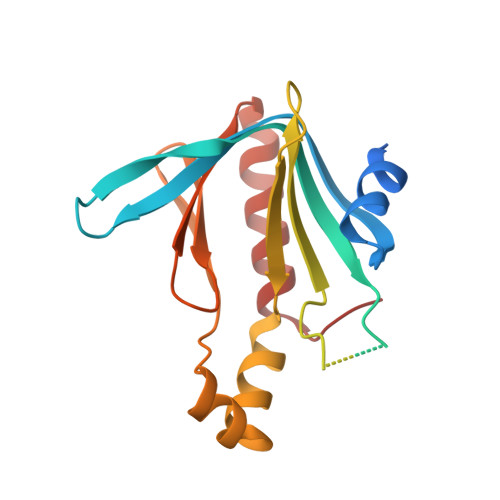
Crystal structure of Dcp1p and its functional implications in mRNA decapping
She, M., Decker, C.J., Sundramurthy, K., Liu, Y., Chen, N., Parker, R., Song, H.(2004) Nat Struct Mol Biol 11: 249-256
- PubMed: 14758354 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb730
- Primary Citation Related Structures:
1Q67 - PubMed Abstract:
A major pathway of eukaryotic mRNA turnover begins with deadenylation, followed by decapping and 5'-->3' exonucleolytic degradation. A critical step in this pathway is decapping, which is carried out by an enzyme composed of Dcp1p and Dcp2p. The crystal structure of Dcp1p shows that it markedly resembles the EVH1 family of protein domains. Comparison of the proline-rich sequence (PRS)-binding sites in this family of proteins with Dcp1p indicates that it belongs to a novel class of EVH1 domains. Mapping of the sequence conservation on the molecular surface of Dcp1p reveals two prominent sites. One of these is required for the function of the Dcp1p-Dcp2p complex, and the other, corresponding to the PRS-binding site of EVH1 domains, is probably a binding site for decapping regulatory proteins. Moreover, a conserved hydrophobic patch is shown to be critical for decapping.
- Laboratory of Macromolecular Structure, Institute of Molecular and Cell Biology, 30 Medical Drive, Singapore 117609.
Organizational Affiliation: