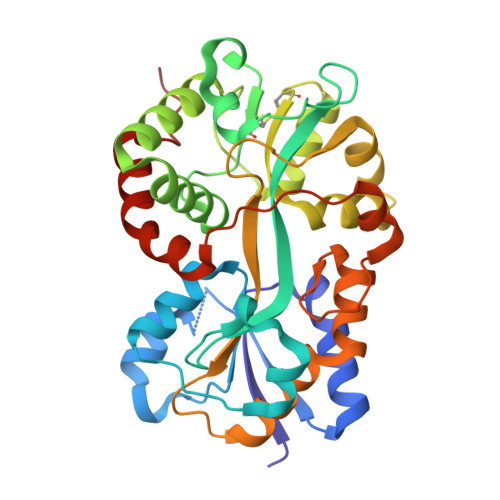
Crystal structure of Pasteurella haemolytica ferric ion-binding protein A reveals a novel class of bacterial iron-binding proteins
Shouldice, S.R., Dougan, D.R., Williams, P.A., Skene, R.J., Snell, G., Scheibe, D., Kirby, S., Hosfield, D.J., McRee, D.E., Schryvers, A.B., Tari, L.W.(2003) J Biological Chem 278: 41093-41098
- PubMed: 12882966 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M306821200
- Primary Citation Related Structures:
1Q35 - PubMed Abstract:
Pasteurellosis caused by the Gram-negative pathogen Pasteurella haemolytica is a serious disease leading to death in cattle. To scavenge growth-limiting iron from the host, the pathogen utilizes the periplasmic ferric ion-binding protein A (PhFbpA) as a component of an ATP-binding cassette transport pathway. We report the 1.2-A structure of the iron-free (apo) form of PhFbpA, which is a member of the transferrin structural superfamily. The protein structure adopts a closed conformation, allowing us to reliably assign putative iron-coordinating residues. Based on our analysis, PhFbpA utilizes a unique constellation of binding site residues and anions to octahedrally coordinate an iron atom. A surprising finding in the structure is the presence of two formate anions on opposite sides of the iron-binding pocket. The formate ions tether the N- and C-terminal domains of the protein and stabilize the closed structure, also providing clues as to probable candidates for synergistic anions in the iron-loaded state. PhFbpA represents a new class of bacterial iron-binding proteins.
- Department of Microbiology and Infectious Diseases, University of Calgary, Calgary, Alberta T2N 4N1, Canada.
Organizational Affiliation: