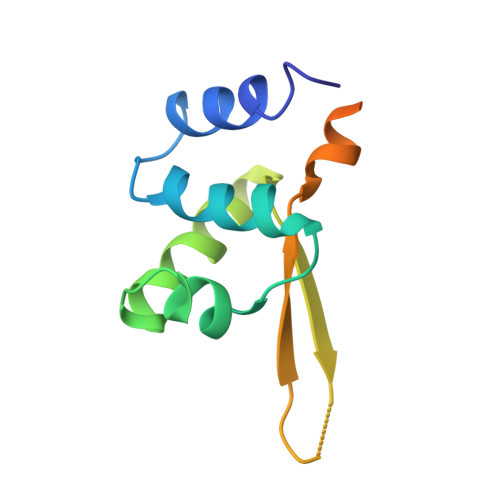
An Extended Winged Helix Domain in General Transcription Factor E/IIE alpha
Meinhart, A., Blobel, J., Cramer, P.(2003) J Biological Chem 278: 48267-48274
- PubMed: 13679366
- DOI: https://doi.org/10.1074/jbc.M307874200
- Primary Citation of Related Structures:
1Q1H - PubMed Abstract:
Initiation of eukaryotic mRNA transcription requires melting of promoter DNA with the help of the general transcription factors TFIIE and TFIIH. Here we define a conserved and functionally essential N-terminal domain in TFE, the archaeal homolog of the large TFIIE subunit alpha. X-ray crystallography shows that this TFE domain adopts a winged helix-turn-helix (winged helix) fold, extended by specific alpha-helices at the N and C termini. Although the winged helix fold is often found in DNA-binding proteins, we show that TFE is not a typical DNA-binding winged helix protein, because its putative DNA-binding face shows a negatively charged groove and an unusually long wing, and because the domain lacks DNA-binding activity in vitro. The groove and a conserved hydrophobic surface patch on the additional N-terminal alpha-helix may, however, allow for interactions with other general transcription factors and RNA polymerase. Homology modeling shows that the TFE domain is conserved in TFIIE alpha, including the potential functional surfaces.
- Institute of Biochemistry, Gene Center, University of Munich, Feodor-Lynen-Strasse 25, 81377 Munich, Germany.
Organizational Affiliation: