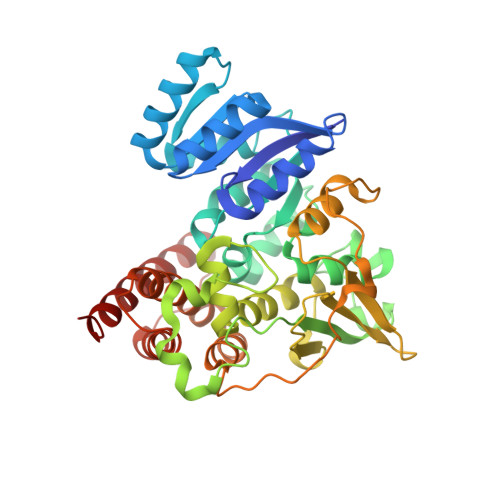
The crystal structure of E.coli 1-deoxy-D-xylulose-5-phosphate reductoisomerase in a ternary complex with the antimalarial compound fosmidomycin and NADPH reveals a tight-binding closed enzyme conformation.
Mac Sweeney, A., Lange, R., Fernandes, R.P., Schulz, H., Dale, G.E., Douangamath, A., Proteau, P.J., Oefner, C.(2005) J Mol Biology 345: 115-127
- PubMed: 15567415 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.10.030
- Primary Citation Related Structures:
1Q0H, 1Q0L, 1Q0Q - PubMed Abstract:
The key enzyme in the non-mevalonate pathway of isoprenoid biosynthesis, 1-deoxy-D-xylulose 5-phosphate reductoisomerase (DXR) has been shown to be the target enzyme of fosmidomycin, an antimalarial, antibacterial and herbicidal compound. Here we report the crystal structure of selenomethionine-labelled Escherichia coli DXR in a ternary complex with NADPH and fosmidomycin at 2.2 A resolution. The structure reveals a considerable conformational rearrangement upon fosmidomycin binding and provides insights into the slow, tight binding inhibition mode of the inhibitor. Although the inhibitor displays an unusual non-metal mediated mode of inhibition, which is an artefact most likely due to the low metal affinity of DXR at the pH used for crystallization, the structural data add valuable information for the rational design of novel DXR inhibitors. Using this structure together with the published structural data and the 1.9 A crystal structure of DXR in a ternary complex with NADPH and the substrate 1-deoxy-D-xylulose 5-phosphate, a model for the physiologically relevant tight-binding mode of inhibition is proposed. The structure of the substrate complex must be interpreted with caution due to the presence of a second diastereomer in the active site.
- Morphochem AG, WRO-1055/338, Schwarzwaldallee 215, CH-4058, Basel, Switzerland.
Organizational Affiliation: