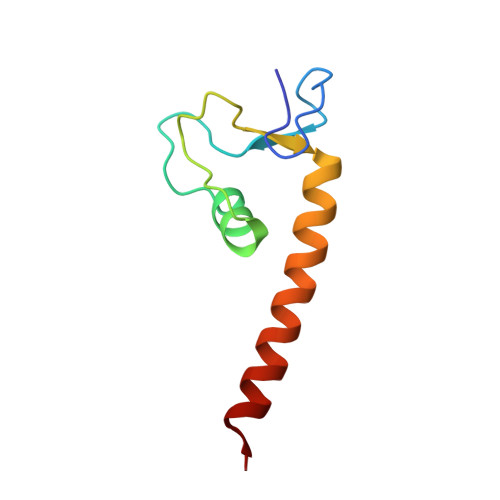
The zinc finger-associated domain of the Drosophila transcription factor grauzone is a novel zinc-coordinating protein-protein interaction module
Jauch, R., Bourenkov, G.P., Chung, H.-R., Urlaub, H., Reidt, U., Wahl, M.C.(2003) Structure 11: 1393-1402
- PubMed: 14604529
- DOI: https://doi.org/10.1016/j.str.2003.09.015
- Primary Citation of Related Structures:
1PZW - PubMed Abstract:
About one-third of the more than 300 C2H2 zinc finger proteins of Drosophila contain a conserved sequence motif, the zinc finger-associated domain (ZAD). Genes that encode ZAD proteins are specific for and expanded in the genomes of insects. Only three ZAD-encoding gene functions are established, and the role of ZAD is unknown. Here we present the crystal structure of the ZAD of Grauzone (ZAD(Grau)), a Drosophila transcription factor that specifically controls the maternal Cdc20-like APC subunit Cortex. ZAD forms an atypical treble-clef-like zinc-coordinating fold. Head-to-tail arrangement of two ZAD(Grau) molecules in the crystals suggests dimer formation, an observation supported by crosslinking and dynamic light scattering. The results indicate that ZAD provides a novel protein-protein interaction module that characterizes a large family of insect transcription factors.
- Max-Planck Institut für biophysikalische Chemie, Abteilung Molekulare Entwicklungsbiologie, Röntgenkristallographie, Am Fassberg 11, D-37077 Göttingen, Germany.
Organizational Affiliation: