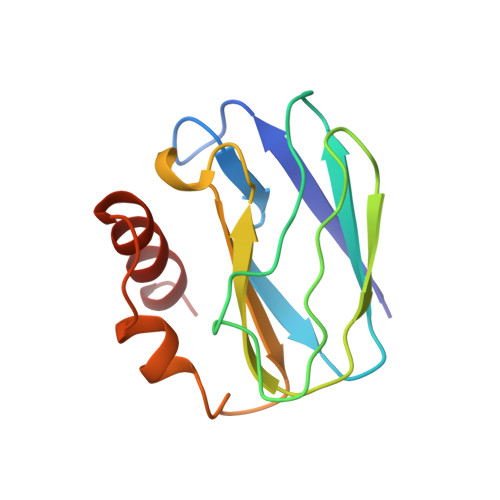
The crystal structure of apo-pseudoazurin from Alcaligenes faecalis S-6.
Petratos, K., Papadovasilaki, M., Dauter, Z.(1995) FEBS Lett 368: 432-434
- PubMed: 7635192 Search on PubMed
- DOI: https://doi.org/10.1016/0014-5793(95)00705-e
- Primary Citation Related Structures:
1PZC - PubMed Abstract:
The 3D structure of the apo-pseudoazurin (copper free pseudoazurin) from Alcaligenes faecalis strain S-6 is determined and refined at pH 6.7 using X-ray diffraction data to 1.85 A resolution. The final crystallographic R-factor is 0.164. Comparing the structures of apo-pseudoazurin and the native (Cu2+) protein, we observed limited differences ranging between 0.1-0.4 A at the vicinity of the copper site, at the loops connecting the secondary structural elements, at certain beta-strands and at the amino and carboxy termini of the protein.
- Institute of Molecular Biology and Biotechnology, Heraklion, Greece.
Organizational Affiliation: