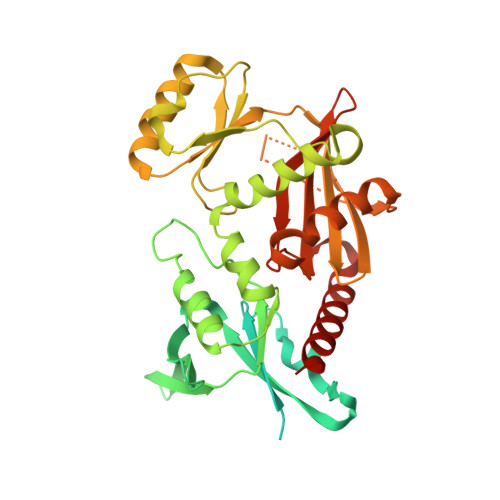
Tetramerization and ATP binding by a protein comprising the A, B, and C domains of rat synapsin I.
Brautigam, C.A., Chelliah, Y., Deisenhofer, J.(2004) J Biological Chem 279: 11948-11956
- PubMed: 14688264 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M312015200
- Primary Citation Related Structures:
1PK8, 1PX2 - PubMed Abstract:
Synapsins are multidomain proteins that are critical for regulating neurotransmitter release in vertebrates. In the present study, two crystal structures of the C domain of rat synapsin I (rSynI-C) in complex with Ca(2+) and ATP reveal that this protein can form a tetramer and that a flexible loop (the "multifunctional loop") contacts bound ATP. Further experiments were carried out on a protein comprising the A, B, and C domains of rat synapsin I (rSynI-ABC). An ATP-stabilized tetramer of rSynI-ABC is observed during velocity sedimentation and size-exclusion chromatographic experiments. These hydrodynamic results also indicate that the A and B domains exist in an extended conformation. Calorimetric measurements of ATP binding to wild-type and mutant rSynI-ABC demonstrate that the multifunctional loop and a cross-tetramer contact are important for ATP binding. The evidence supports a view of synapsin I as an ATP-utilizing, tetrameric protein made up of monomers that have a flexible, extended N terminus.
- Howard Hughes Medical Institute and the Department of Biochemistry, The University of Texas Southwestern Medical Center at Dallas, Dallas, Texas 75390-9050, USA.
Organizational Affiliation: