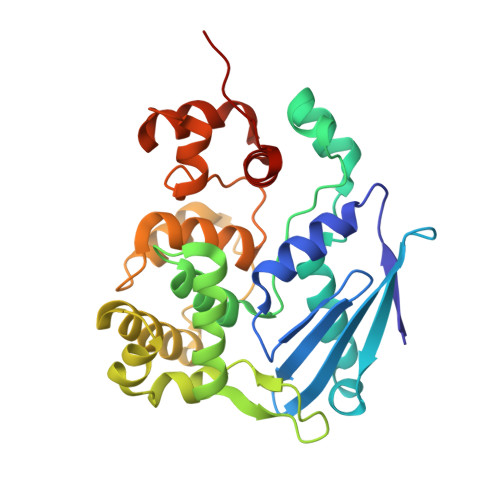
3-methyladenine-DNA glycosylase II: the crystal structure of an AlkA-hypoxanthine complex suggests the possibility of product inhibition.
Teale, M., Symersky, J., DeLucas, L.(2002) Bioconjug Chem 13: 403-407
- PubMed: 12009927
- DOI: https://doi.org/10.1021/bc015527v
- Primary Citation Related Structures:
1PVS - PubMed Abstract:
Escherichia coli (E. coli) protein 3-methyladenine-DNA glycosylase II (AlkA) functions primarily by removing alkylation damage from duplex and single stranded DNA. A crystal structure of AlkA was refined to 2.0 A resolution. This structure in turn was used to refine an AlkA-hypoxanthine (substrate) complex structure to 2.4 A resolution. The complex structure shows hypoxanthine located in AlkA's active site stacked between residues W218 and Y239. The structural analysis of the AlkA and AlkA-hypoxanthine structures indicate that free hypoxanthine binding in the active site may inhibit glycosylase activity.
- Laboratory for Structural Biology, University of Alabama in Huntsville, 35899, USA. tealem@email.uah.edu
Organizational Affiliation: