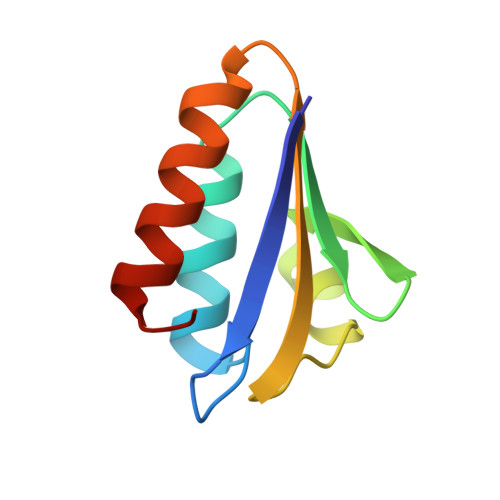
The 1.6 A structure of histidine-containing phosphotransfer protein HPr from Streptococcus faecalis.
Jia, Z., Vandonselaar, M., Hengstenberg, W., Quail, J.W., Delbaere, L.T.(1994) J Mol Biology 236: 1341-1355
- PubMed: 8126724 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(94)90062-0
- Primary Citation Related Structures:
1PTF - PubMed Abstract:
The histidine-containing phosphocarrier protein (HPr) is a central component of the phosphoenolpyruvate: sugar phosphotransferase system (PTS) that transports carbohydrates across the cell membrane of bacteria. The three-dimensional structure of Gram-positive Streptococcus faecalis HPr has been determined using the method of multiple isomorphous replacement. The R factor for all data is 0.156 for S. faecalis HPr at 1.6 A resolution with very good geometry. The overall folding topology of HPr is a classical open-faced beta-sandwich, consisting of four antiparallel beta-strands and three alpha-helices. Remarkable disallowed Ramachandran torsion angles of Ala16 at the active center, revealed by the X-ray structure of S. faecalis HPr, demonstrate a unique example of torsion-angle strain that is likely involved directly in protein function. A brief report concerning the torsion-angle strain has been presented recently. A newly-determined pH 7.0 structure is shown to have the same open conformation of the active center and the same torsion-angle strain at Ala16, suggesting that pH is not responsible for the structural observations. The current structure suggests a role for residues 12 and 51 in HPr's function, since they are involved in the active center through direct and indirect hydrogen-bonding interactions with the imidazole ring of His15. It is found that Ser46, the regulatory site in HPr from Gram-positive bacteria, N-caps the minor alpha-B helix and is also involved in the Asn43-Ser46 beta-turn. This finding, in conjunction with the proposed routes of communication between the regulatory site Ser46 and the active center in S. faecalis HPr, provides new insight into the understanding of how Ser46 might function. The putative involvement of the C-terminal alpha-carboxyl group and the related Gly67-Glu70 reverse beta-turn with respect to the function of HPr are described.
- Department of Chemistry, University of Saskatchewan, Saskatoon, Canada.
Organizational Affiliation: