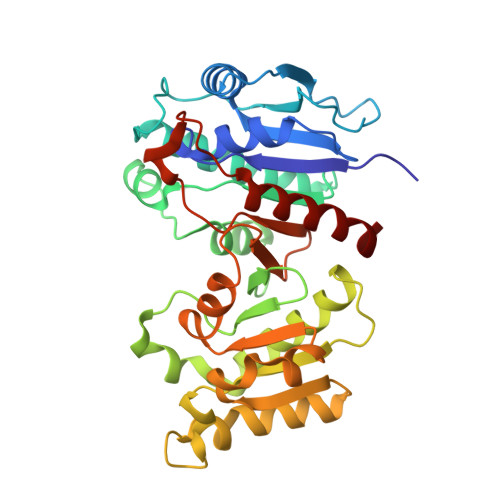
Crystal Structure of Escherichia coli PdxA, an Enzyme Involved in the Pyridoxal Phosphate Biosynthesis Pathway
Sivaraman, J., Li, Y., Banks, J., Cane, D.E., Matte, A., Cygler, M.(2003) J Biological Chem 278: 43682-43690
- PubMed: 12896974 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M306344200
- Primary Citation Related Structures:
1PS6, 1PS7, 1PTM - PubMed Abstract:
Pyridoxal 5'-phosphate is an essential cofactor for many enzymes responsible for the metabolic conversions of amino acids. Two pathways for its de novo synthesis are known. The pathway utilized by Escherichia coli consists of six enzymatic steps catalyzed by six different enzymes. The fourth step is catalyzed by 4-hydroxythreonine-4-phosphate dehydrogenase (PdxA, E.C. 1.1.1.262), which converts 4-hydroxy-l-threonine phosphate (HTP) to 3-amino-2-oxopropyl phosphate. This divalent metal ion-dependent enzyme has a strict requirement for the phosphate ester form of the substrate HTP, but can utilize either NADP+ or NAD+ as redox cofactor. We report the crystal structure of E. coli PdxA and its complex with HTP and Zn2+. The protein forms tightly bound dimers. Each monomer has an alpha/beta/alpha-fold and can be divided into two subdomains. The active site is located at the dimer interface, within a cleft between the two subdomains and involves residues from both monomers. A Zn2+ ion is bound within each active site, coordinated by three conserved histidine residues from both monomers. In addition two conserved amino acids, Asp247 and Asp267, play a role in maintaining integrity of the active site. The substrate is anchored to the enzyme by the interactions of its phospho group and by coordination of the amino and hydroxyl groups by the Zn2+ ion. PdxA is structurally similar to, but limited in sequence similarity with isocitrate dehydrogenase and isopropylmalate dehydrogenase. These structural similarities and the comparison with a NADP-bound isocitrate dehydrogenase suggest that the cofactor binding mode of PdxA is very similar to that of the other two enzymes and that PdxA catalyzes a stepwise oxidative decarboxylation of the substrate HTP.
- Biotechnology Research Institute, NRC, Montreal, Quebec, Canada.
Organizational Affiliation: