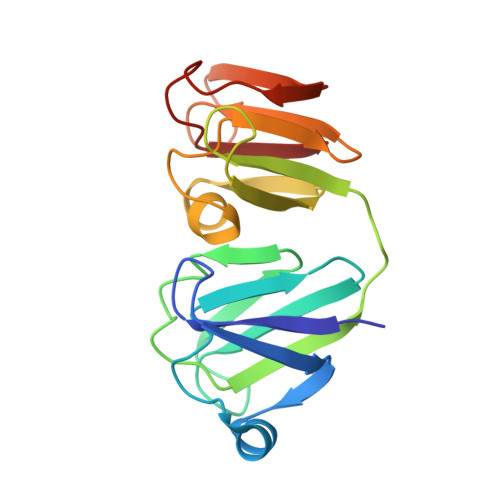
NMR-derived three-dimensional solution structure of protein S complexed with calcium.
Bagby, S., Harvey, T.S., Eagle, S.G., Inouye, S., Ikura, M.(1994) Structure 2: 107-122
- PubMed: 8081742
- DOI: https://doi.org/10.1016/s0969-2126(00)00013-7
- Primary Citation Related Structures:
1PRR, 1PRS - PubMed Abstract:
Protein S is a developmentally-regulated Ca(2+)-binding protein of the soil bacterium Myxococcus xanthus. It functions by forming protective, multilayer spore surface assemblies which may additionally act as a cell-cell adhesive. Protein S is evolutionarily related to vertebrate lens beta gamma-crystallins. The three-dimensional solution structure of Ca(2+)-loaded protein S has been determined using multi-dimensional heteronuclear NMR spectroscopy. (Sixty structures were calculated, from which thirty were selected with a root mean square difference from the mean of 0.38 A for backbone atoms and 1.22 A for all non-hydrogen atoms.) The structure was analyzed and compared in detail with X-ray crystallographic structures of beta gamma-crystallins. The two internally homologous domains of protein S were compared, and hydrophobic cores, domain interfaces, surface ion pairing, amino-aromatic interactions and potential modes of multimerization are discussed. Structural features of protein S described here help to explain its overall thermostability, as well as the higher stability and Ca2+ affinity of the amino-terminal domain relative to the carboxy-terminal domain. Two potential modes of multimerization are proposed involving cross-linking of protein S molecules through surface Ca(2+)-binding sites and formation of the intramolecular protein S or gamma B-crystallin interdomain interface in an intermolecular content. This structural analysis may also have implications for Ca(2+)-dependent cell-cell interactions mediated by the vertebrate cadherins and Dictyostelium discoideum protein gp24.
- Division of Molecular and Structural Biology, Ontario Cancer Institute, Toronto, Canada.
Organizational Affiliation: