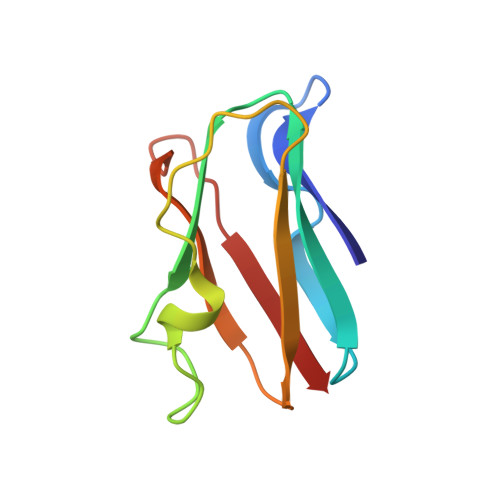
High-resolution solution structure of reduced parsley plastocyanin.
Bagby, S., Driscoll, P.C., Harvey, T.S., Hill, H.A.(1994) Biochemistry 33: 6611-6622
- PubMed: 8204598 Search on PubMed
- DOI: https://doi.org/10.1021/bi00187a031
- Primary Citation Related Structures:
1PLA, 1PLB - PubMed Abstract:
A high-resolution three-dimensional solution structure of parsley plastocyanin has been determined using 1H-NMR-derived data. An ensemble of 30 conformers has been calculated, exhibiting an atomic root mean square distribution about the mean coordinate positions of 0.37 +/- 0.03 A for backbone atoms and 0.75 +/- 0.04 A for all heavy atoms. (These values exclude residues 8-10 which are disordered.) The global fold of parsley plastocyanin is closely similar to those of other plastocyanins which have been structurally characterized by X-ray diffraction and NMR methods. However, deletion of residues at positions 57 and 58 of the consensus plastocyanin sequence causes elimination of a turn found in most higher plant plastocyanins. This turn is located in an acidic patch binding site, which consists of two clusters of acidic residues at positions 42-45 and 59-61. These residues surround the side chain of Tyr 83, which has been shown to be involved in binding of and electron transfer from cytochrome f, one of plastocyanin's physiological partners. The acidic recognition site is further disrupted in parsley plastocyanin by nonconservative substitution of two charged residues at positions 59 and 60. The NMR-derived structures show that E53, E85, and E95 compensate for these substitutions and give parsley plastocyanin an acidic recognition site of similar extent to that of other higher plant plastocyanins.(ABSTRACT TRUNCATED AT 250 WORDS)