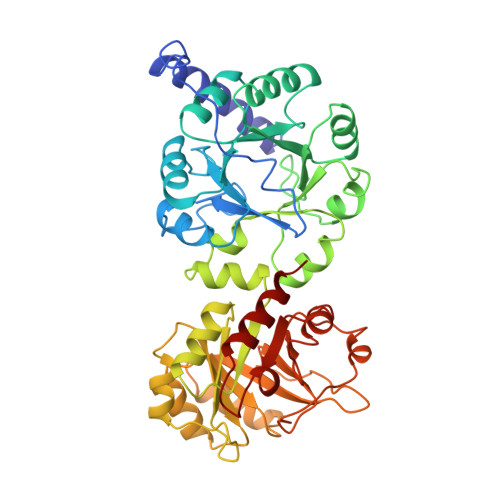
Three-dimensional structure of the bifunctional enzyme phosphoribosylanthranilate isomerase: indoleglycerolphosphate synthase from Escherichia coli refined at 2.0 A resolution.
Wilmanns, M., Priestle, J.P., Niermann, T., Jansonius, J.N.(1992) J Mol Biology 223: 477-507
- PubMed: 1738159 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(92)90665-7
- Primary Citation Related Structures:
1PII - PubMed Abstract:
The three-dimensional structure of the monomeric bifunctional enzyme N-(5'-phosphoribosyl)anthranilate isomerase:indole-3-glycerol-phosphate synthase from Escherichia coli has been refined at 2.0 A resolution, using oscillation film data obtained from synchrotron radiation. The model includes the complete protein (452 residues), two phosphate ions and 628 water molecules. The final R-factor is 17.3% for all observed data between 15 and 2 A resolution. The root-mean-square deviations from ideal bond lengths and bond angles are 0.010 A and 3.2 degrees, respectively. The structure of N-(5'-phosphoribosyl)anthranilate isomerase: indole-3-glycerol-phosphate synthase from E. coli comprises two beta/alpha-barrel domains that superimpose with a root-mean-square deviation of 2.03 A for 138 C alpha-pairs. The C-terminal domain (residues 256 to 452) catalyses the PRAI reaction and the N-terminal domain (residues 1 to 255) catalyses the IGPS reaction, two sequential steps in tryptophan biosynthesis. The enzyme has the overall shape of a dumb-bell, resulting in a surface area that is considerably larger than normally observed for monomeric proteins of this size. The active sites of the PRAI and the IGPS domains, both located at the C-terminal side of the central beta-barrel, contain equivalent binding sites for the phosphate moieties of the substrates N-(5'-phosphoribosyl) anthranilate and 1-(o-carboxyphenylamino)-1-deoxyribulose-5-phosphate. These two phosphate binding sites are identical with respect to their positions within the tertiary structure of the beta/alpha-barrel, the conformation of the residues involved in phosphate binding and the hydrogen-bonding network between the phosphate ions and the protein. The active site cavities of both domains contain similar hydrophobic pockets that presumably bind the anthranilic acid moieties of the substrates. These similarities of the tertiary structures and the active sites of the two domains provide evidence that N-(5'-phosphoribosyl)anthranilate isomerase:indole-3-glycerol-phosphate synthase from E. coli results from a gene duplication event of a monomeric beta/alpha-barrel ancestor.
- Department of Structural Biology, University of Basel, Switzerland.
Organizational Affiliation: