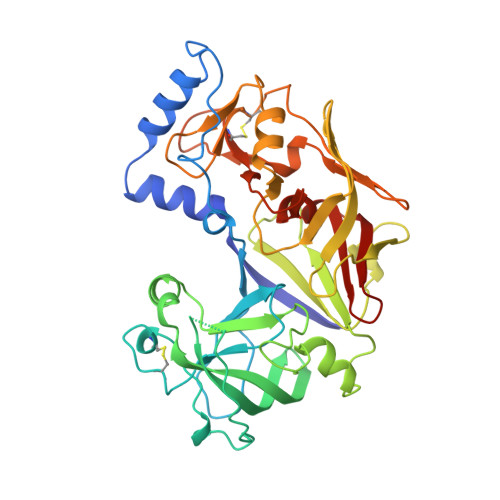
Crystal structure of the novel aspartic proteinase zymogen proplasmepsin II from plasmodium falciparum.
Bernstein, N.K., Cherney, M.M., Loetscher, H., Ridley, R.G., James, M.N.(1999) Nat Struct Biol 6: 32-37
- PubMed: 9886289 Search on PubMed
- DOI: https://doi.org/10.1038/4905
- Primary Citation Related Structures:
1PFZ - PubMed Abstract:
Proplasmepsin II is the zymogen of plasmepsin II, an aspartic proteinase used by Plasmodiumfalciparum to digest hemoglobin during the blood stage of malaria. A large shift between the N-domain and the central and C-domains of proplasmepsin II opens the active site cleft, preventing the formation of a functional aspartic proteinase active site. This mode of inhibition of catalytic activity has not been observed in any other aspartic proteinase zymogen. Instead of occluding a pre-formed active site, as in the gastric aspartic proteinase zymogens, the prosegment of proplasmepsin II interacts extensively with the C-domain and serves as a 'harness' to keep the domains apart. Disruption of key salt bridges at low pH may be important in activation.
- Medical Research Council Group in Protein Structure and Function, Department of Biochemistry, University of Alberta, Edmonton, Canada.
Organizational Affiliation: