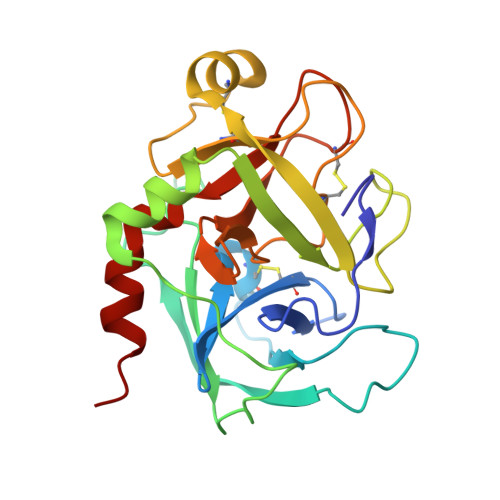
X-ray structure of clotting factor IXa: active site and module structure related to Xase activity and hemophilia B.
Brandstetter, H., Bauer, M., Huber, R., Lollar, P., Bode, W.(1995) Proc Natl Acad Sci U S A 92: 9796-9800
- PubMed: 7568220 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.92.21.9796
- Primary Citation Related Structures:
1PFX - PubMed Abstract:
Hereditary deficiency of factor IXa (fIXa), a key enzyme in blood coagulation, causes hemophilia B, a severe X chromosome-linked bleeding disorder afflicting 1 in 30,000 males; clinical studies have identified nearly 500 deleterious variants. The x-ray structure of porcine fIXa described here shows the atomic origins of the disease, while the spatial distribution of mutation sites suggests a structural model for factor X activation by phospholipid-bound fIXa and cofactor VIIIa. The 3.0-A-resolution diffraction data clearly show the structures of the serine proteinase module and the two preceding epidermal growth factor (EGF)-like modules; the N-terminal Gla module is partially disordered. The catalytic module, with covalent inhibitor D-Phe-1I-Pro-2I-Arg-3I chloromethyl ketone, most closely resembles fXa but differs significantly at several positions. Particularly noteworthy is the strained conformation of Glu-388, a residue strictly conserved in known fIXa sequences but conserved as Gly among other trypsin-like serine proteinases. Flexibility apparent in electron density together with modeling studies suggests that this may cause incomplete active site formation, even after zymogen, and hence the low catalytic activity of fIXa. The principal axes of the oblong EGF-like domains define an angle of 110 degrees, stabilized by a strictly conserved and fIX-specific interdomain salt bridge. The disorder of the Gla module, whose hydrophobic helix is apparent in electron density, can be attributed to the absence of calcium in the crystals; we have modeled the Gla module in its calcium form by using prothrombin fragment 1. The arched module arrangement agrees with fluorescence energy transfer experiments. Most hemophilic mutation sites of surface fIX residues occur on the concave surface of the bent molecule and suggest a plausible model for the membrane-bound ternary fIXa-FVIIIa-fX complex structure: fIXa and an equivalently arranged fX arch across an underlying fVIIIa subdomain from opposite sides; the stabilizing fVIIIa interactions force the catalytic modules together, completing fIXa active site formation and catalytic enhancement.
- Max-Planck-Institut für Biochemie, Martinsried, Germany.
Organizational Affiliation: