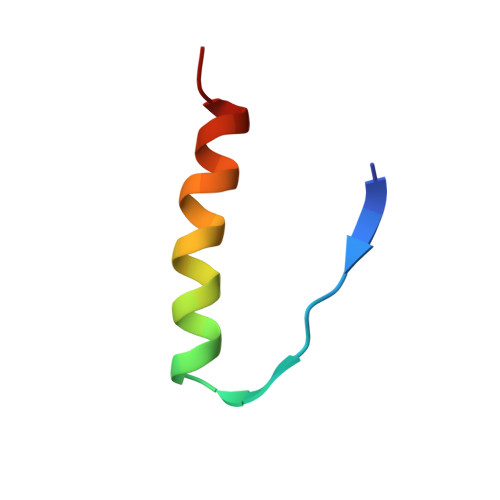
Solution structure of the tetrameric minimum transforming domain of p53.
Lee, W., Harvey, T.S., Yin, Y., Yau, P., Litchfield, D., Arrowsmith, C.H.(1994) Nat Struct Biol 1: 877-890
- PubMed: 7773777 Search on PubMed
- DOI: https://doi.org/10.1038/nsb1294-877
- Primary Citation Related Structures:
1PES, 1PET - PubMed Abstract:
We report the solution structure of the minimum transforming domain (residues 303-366) of human p53 (p53tet) determined by multidimensional NMR spectroscopy. This domain contains a number of important functions associated with p53 activity including transformation, oligomerization, nuclear localization and a phosphorylation site for p34/cdc2 kinase. p53tet forms a symmetric dimer of dimers that is significantly different from a recent structure reported for a shorter construct of this domain. Phosphorylation of Ser 315 has only minor structural consequences, as this region of the protein is unstructured. Modelling based on the p53tet structure suggests possible modes of interaction between adjacent domains in full-length p53 as well as modes of interaction with DNA.
- Division of Molecular and Structural Biology, Ontario Cancer Institute, Toronto, Canada.
Organizational Affiliation: