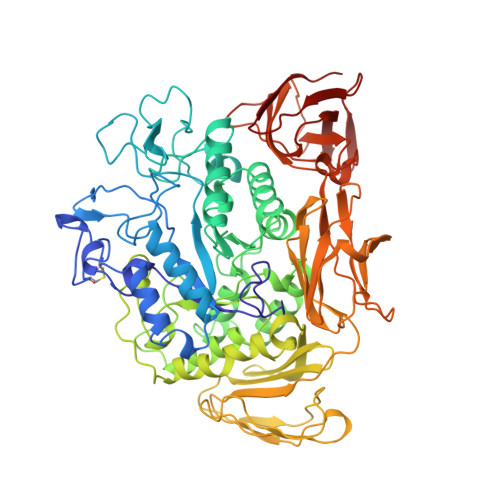
X-ray structure of cyclodextrin glucanotransferase from alkalophilic Bacillus sp. 1011. Comparison of two independent molecules at 1.8 A resolution.
Harata, K., Haga, K., Nakamura, A., Aoyagi, M., Yamane, K.(1996) Acta Crystallogr D Biol Crystallogr 52: 1136-1145
- PubMed: 15299574 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444996008438
- Primary Citation Related Structures:
1PAM - PubMed Abstract:
Cyclodextrin glucanotransferase (CGTase) is an enzyme which produces cyclodextrins by the degradation of starch. The enzyme from alkalophilic Bacillus sp. 1011, consisting of 686 amino acid residues, was crystallized from the solution containing 20% PEG 3000 and 20% 2-propanol at pH 5.6 adjusted with citrate buffer. The space group was P1 and the unit cell contained two molecules (V(m) = 2.41 A(3) Da(-1)). The structure was solved by the molecular replacement method and refined to a conventional R value of 0.161 (R(free) = 0.211) for the reflections in the resolution range 1.8-10 A by energy minimization combined with simulated annealing. The molecule consists of five domains, designated A-E, and its backbone structure is similar to the structure of other bacterial CGTases. The molecule has two calcium binding sites where calcium ions are coordinated by seven ligands, forming a distorted pentagonal bipyramid. The two independent molecules are related by a pseudotwofold symmetry and are superimposed with an r.m.s. deviation value of 0.32 A for equivalent C(alpha) atoms. Comparison of these molecules indicated the relatively large mobility of domains C and E with respect to domain A. The active site is filled with water molecules forming a hydrogen-bond network with polar side-chain groups. Two water molecules commonly found in the active center of both molecules link to several catalytically important residues by hydrogen bonds and participate in maintaining a similar orientation of side chains in the two independent molecules.
- Biomolecules Department, National Institute of Bioscience and Human Technology, Tsukuba, Ibaraki, Japan. harata@nibh.go.jp
Organizational Affiliation: