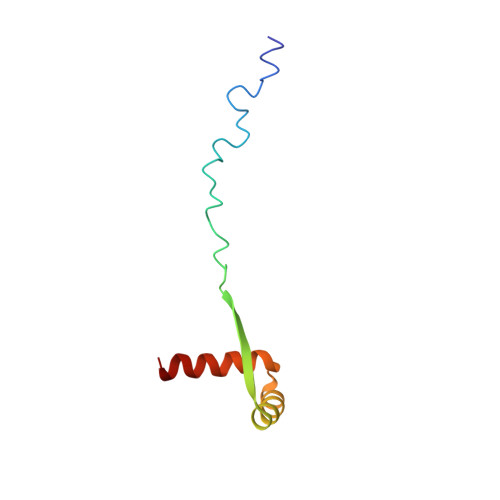
ParG, a protein required for active partition of bacterial plasmids, has a dimeric ribbon-helix-helix structure.
Golovanov, A.P., Barilla, D., Golovanova, M., Hayes, F., Lian, L.Y.(2003) Mol Microbiol 50: 1141-1153
- PubMed: 14622405
- DOI: https://doi.org/10.1046/j.1365-2958.2003.03750.x
- Primary Citation Related Structures:
1P94 - PubMed Abstract:
The ParG protein (8.6 kDa) is an essential component of the DNA partition complex of multidrug resistance plasmid TP228. ParG is a dimer in solution, interacts with DNA sequences upstream of the parFG genes and also with the ParF partition protein both in the absence and presence of target DNA. Here, the solution nuclear magnetic resonance structure of ParG is reported. The ParG dimer is composed of a folded domain formed by two closely intertwined C-terminal parts (residues 33-76), and two highly mobile tails consisting of N-terminal regions (residues 1-32). The folded part of ParG has the ribbon-helix-helix (RHH) architecture similar to that of the Arc/MetJ superfamily of DNA-binding transcriptional repressors, although the primary sequence similarity is very low. ParG interacts with DNA predominantly via its folded domain; this interaction is coupled with ParG oligomerization. The dimeric RHH structure of ParG suggests that it binds to DNA by inserting the double-stranded beta-sheet into the major groove of DNA, in a manner similar to transcriptional repressors from the Arc/MetJ superfamily, and that ParG can function as a transcriptional repressor itself. A new classification of proteins belonging to the Arc/MetJ superfamily and ParG homologues is proposed, based on the location of a conserved positively charged residue at either the beginning or at the end of the beta-strand which forms part of the DNA recognition motif.
- Department of Biomolecular Sciences, University of Manchester Institute of Science and Technology (UMIST), PO Box 88, Manchester M60 1QD, UK.
Organizational Affiliation: