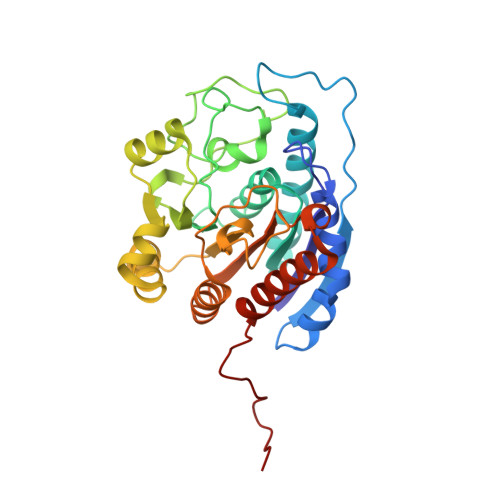
Structural and functional importance of first-shell metal ligands in the binuclear manganese cluster of arginase I
Cama, E., Emig, F.A., Ash, D.E., Christianson, D.W.(2003) Biochemistry 42: 7748-7758
- PubMed: 12820884 Search on PubMed
- DOI: https://doi.org/10.1021/bi030074y
- Primary Citation Related Structures:
1P8M, 1P8N, 1P8O, 1P8P, 1P8Q, 1P8R, 1P8S - PubMed Abstract:
Arginase is a binuclear manganese metalloenzyme that hydrolyzes l-arginine to form l-ornithine and urea. The three-dimensional structures of D128E, D128N, D232A, D232C, D234E, H101N, and H101E arginases I have been determined by X-ray crystallographic methods to elucidate the roles of the first-shell metal ligands in the stability and catalytic activity of the enzyme. This work represents the first structure-based dissection of the binuclear manganese cluster using site-directed mutagenesis and X-ray crystallography. Substitution of the metal ligands compromises the catalytic activity of the enzyme, either by the loss or disruption of the metal cluster or the nucleophilic metal-bridging hydroxide ion. However, the substitution of the metal ligands or the reduction of Mn(2+)(A) or Mn(2+)(B) occupancy does not compromise enzyme-substrate affinity as reflected by K(M), which remains relatively invariant across this series of arginase variants. This implicates a nonmetal binding site for substrate l-arginine in the precatalytic Michaelis complex, as proposed based on analysis of the native enzyme structure (Kanyo, Z. F., Scolnick, L. R., Ash, D. E., and Christianson, D. W. (1996) Nature 383, 554-557).
- Roy and Diana Vagelos Laboratories, Department of Chemistry, University of Pennsylvania, Philadelphia, Pennsylvania 19104-6323, USA.
Organizational Affiliation: