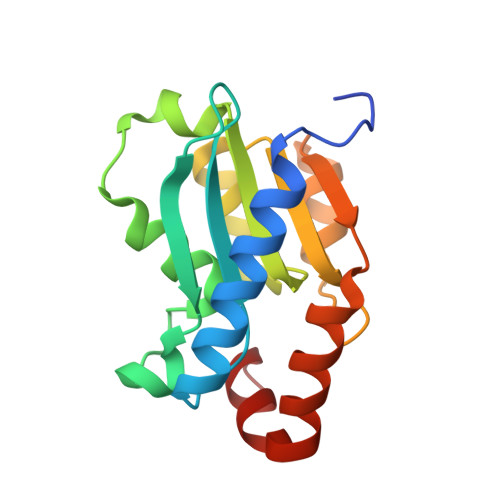
The 1.14 a crystal structure of yeast Cytosine deaminase. Evolution of nucleotide salvage enzymes and implications for genetic chemotherapy.
Ireton, G.C., Black, M.E., Stoddard, B.L.(2003) Structure 11: 961-972
- PubMed: 12906827
- DOI: https://doi.org/10.1016/s0969-2126(03)00153-9
- Primary Citation of Related Structures:
1OX7, 1P6O - PubMed Abstract:
Cytosine deaminase (CD) catalyzes the deamination of cytosine and is only present in prokaryotes and fungi, where it is a member of the pyrimidine salvage pathway. The enzyme is of interest both for antimicrobial drug design and gene therapy applications against tumors. The structure of Saccharomyces cerevisiae CD has been determined in the presence and absence of a mechanism-based inhibitor, at 1.14 and 1.43 A resolution, respectively. The enzyme forms an alpha/beta fold similar to bacterial cytidine deaminase, but with no similarity to the alpha/beta barrel fold used by bacterial cytosine deaminase or mammalian adenosine deaminase. The structures observed for bacterial, fungal, and mammalian nucleic acid deaminases represent an example of the parallel evolution of two unique protein folds to carry out the same reaction on a diverse array of substrates.
- Fred Hutchinson Cancer Research Center, University of Washington, 1100 Fairview Avenue North A3-023, Seattle, WA 98109, USA.
Organizational Affiliation: