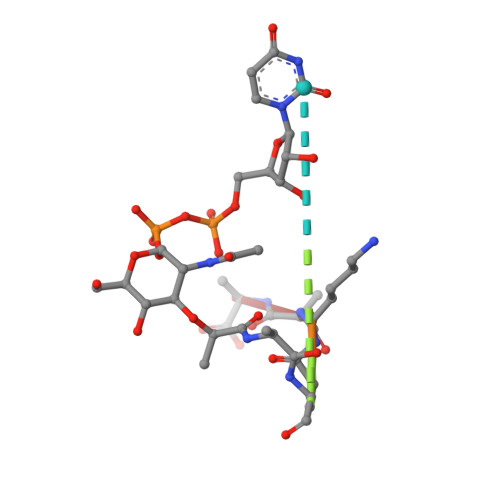
Crystal structures of Weissella viridescens FemX and its complex with UDP-MurNAc-pentapeptide: insights into FemABX family substrates recognition.
Biarrotte-Sorin, S., Maillard, A.P., Delettre, J., Sougakoff, W., Arthur, M., Mayer, C.(2004) Structure 12: 257-267
- PubMed: 14962386 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.01.006
- Primary Citation Related Structures:
1NE9, 1P4N - PubMed Abstract:
Members of the FemABX protein family are novel therapeutic targets, as they are involved in the synthesis of the bacterial cell wall. They catalyze the addition of amino acid(s) on the peptidoglycan precursor using aminoacylated tRNA as a substrate. We report here the high-resolution structure of Weissella viridescens L-alanine transferase FemX and its complex with the UDP-MurNAc-pentapeptide. This is the first structure example of a FemABX family member that does not possess a coiled-coil domain. FemX consists of two structurally equivalent domains, separated by a cleft containing the binding site of the UDP-MurNAc-pentapeptide and a long channel that traverses one of the two domains. Our structural studies bring new insights into the evolution of the FemABX and the related GNAT superfamilies, shed light on the recognition site of the aminoacylated tRNA in Fem proteins, and allowed manual docking of the acceptor end of the alanyl-tRNAAla.
- Laboratoire de Minéralogie-Cristallographie de Paris, Université Paris 6, 4 place Jussieu, Paris Cedex 05, 75252, France.
Organizational Affiliation: