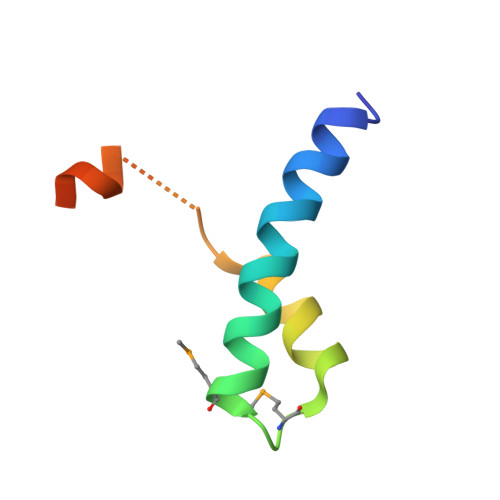
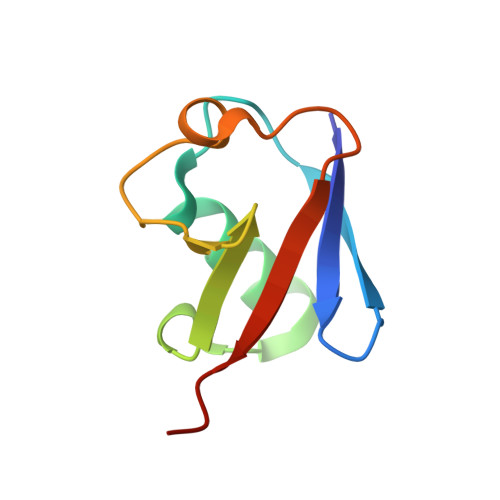
Mechanism of Ubiquitin Recognition by the CUE Domain of Vps9p.
Prag, G., Misra, S., Jones, E.A., Ghirlando, R., Davies, B.A., Horazdovsky, B.F., Hurley, J.H.(2003) Cell 113: 609-620
- PubMed: 12787502
- DOI: https://doi.org/10.1016/s0092-8674(03)00364-7
- Primary Citation Related Structures:
1MN3, 1P3Q - PubMed Abstract:
Coupling of ubiquitin conjugation to ER degradation (CUE) domains are approximately 50 amino acid monoubiquitin binding motifs found in proteins of trafficking and ubiquitination pathways. The 2.3 A structure of the Vps9p-CUE domain is a dimeric domain-swapped variant of the ubiquitin binding UBA domain. The 1.7 A structure of the CUE:ubiquitin complex shows that one CUE dimer binds one ubiquitin molecule. The bound CUE dimer is kinked relative to the unbound CUE dimer and wraps around ubiquitin. The CUE monomer contains two ubiquitin binding surfaces on opposite faces of the molecule that cannot bind simultaneously to a single ubiquitin molecule. Dimerization of the CUE domain allows both surfaces to contact a single ubiquitin molecule, providing a mechanism for high-affinity binding to monoubiquitin.
- Laboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Department of Health and Human Services, Bethesda, MD 20892, USA.
Organizational Affiliation: