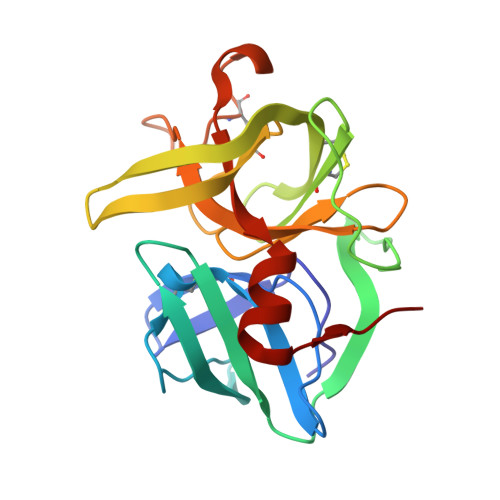
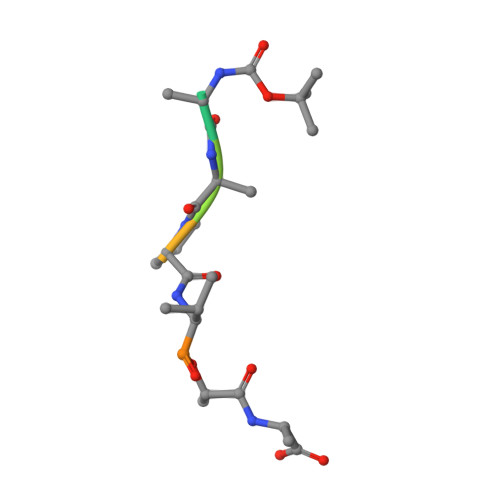
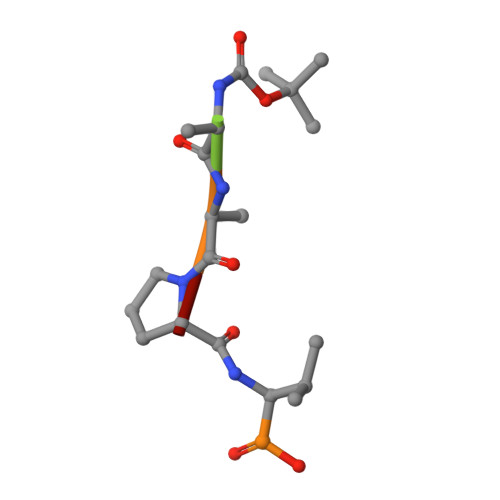
Crystal structures of alpha-lytic protease complexes with irreversibly bound phosphonate esters.
Bone, R., Sampson, N.S., Bartlett, P.A., Agard, D.A.(1991) Biochemistry 30: 2263-2272
- PubMed: 1998685 Search on PubMed
- DOI: https://doi.org/10.1021/bi00222a032
- Primary Citation Related Structures:
1P11, 1P12 - PubMed Abstract:
The structures of the complexes with alpha-lytic protease of both phosphorus stereoisomers of N-[(2S)-2-[[[(1R)-1-[N-[(tert-butyloxycarbonyl)-L-alanyl-L-alanyl- L-prolyl]amino]-2-methylpropyl]-phenoxyphosphinyl]oxy]propanoyl]- L-alanine methyl ester, an analogue of the peptide Boc-Ala-Ala-Pro-Val-Ala-Ala where Val is replaced with an analogous phosphonate phenyl ester and the subsequent Ala is replaced with lactate, have been determined to high resolution (1.9 A) by X-ray crystallography. Both stereoisomers inactivate the enzyme but differ by a factor of 2 in the second-order rate constant for inactivation [Sampson, N. S., & Bartlett, P. A. (1991) Biochemistry (preceding paper in this issue)]. One isomer (B) forms a tetrahedral adduct in which the phosphonate phenyl ester is displaced by the active site serine (S195) and interacts with the enzyme across seven substrate recognition sites that span both sides of the scissile bond. Seven hydrogen bonds are formed with the enzyme, and 510 A2 of hydrophobic surface area is buried when the inhibitor interacts with the enzyme. Although two hydrogen bonds are gained by incorporation of two residues on the C-terminal side of the scissile bond into the inhibitor, there is very little adjustment in the structure of the enzyme in this region. Surprisingly, the active site histidine (H57) does not interact with the phosphonate, apparently because the phosphonate lacks negative charge in or near the oxyanion hole, and instead, the side chain rotates out of the active site cleft and hydrogen bonds with solvent. The other isomer (A) forms a mixture of two different tetrahedral adducts in the active site, both covalently bonded to Ser 195. One adduct, at approximately 58% occupancy, is exactly the same in structure as the complex formed with isomer B, and the other adduct, at 42% occupancy, has lost the two residues C-terminal to the scissile bond by hydrolysis. In the lower occupancy structure, His 57 does not rotate out of the active site and forms a hydrogen bond with the phosphonate oxygen instead. The structures of both complexes were insensitive to pH. As very little change in structure accompanies the histidine rotation, the complex with isomer B provides an excellent mimic for the structure of the transition state (or high-energy reaction intermediate) that spans both sides of the scissile bond.
- Department of Biochemistry and Biophysics, University of California, San Francisco 94143-0448.
Organizational Affiliation: