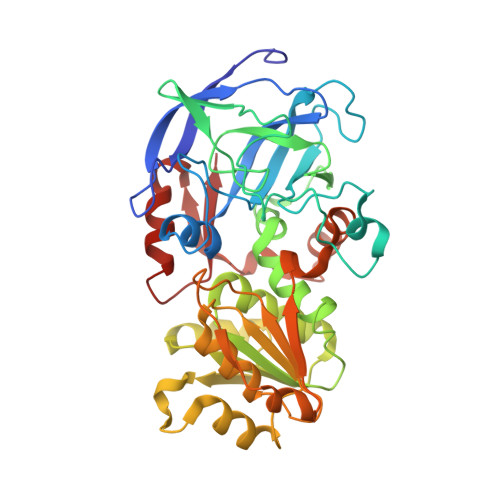
Crystal structure of the vertebrate NADP(H)-dependent alcohol dehydrogenase (ADH8)
Rosell, A., Valencia, E., Pares, X., Fita, I., Farres, J., Ochoa, W.F.(2003) J Mol Biology 330: 75-85
- PubMed: 12818203 Search on PubMed
- DOI: https://doi.org/10.1016/s0022-2836(03)00431-5
- Primary Citation Related Structures:
1P0C, 1P0F - PubMed Abstract:
The amphibian enzyme ADH8, previously named class IV-like, is the only known vertebrate alcohol dehydrogenase (ADH) with specificity towards NADP(H). The three-dimensional structures of ADH8 and of the binary complex ADH8-NADP(+) have been now determined and refined to resolutions of 2.2A and 1.8A, respectively. The coenzyme and substrate specificity of ADH8, that has 50-65% sequence identity with vertebrate NAD(H)-dependent ADHs, suggest a role in aldehyde reduction probably as a retinal reductase. The large volume of the substrate-binding pocket can explain both the high catalytic efficiency of ADH8 with retinoids and the high K(m) value for ethanol. Preference of NADP(H) appears to be achieved by the presence in ADH8 of the triad Gly223-Thr224-His225 and the recruitment of conserved Lys228, which define a binding pocket for the terminal phosphate group of the cofactor. NADP(H) binds to ADH8 in an extended conformation that superimposes well with the NAD(H) molecules found in NAD(H)-dependent ADH complexes. No additional reshaping of the dinucleotide-binding site is observed which explains why NAD(H) can also be used as a cofactor by ADH8. The structural features support the classification of ADH8 as an independent ADH class.
- Departament de Bioquímica i Biologia Molecular, Universitat Autònoma de Barcelona, 08193 Bellaterra, Spain.
Organizational Affiliation: