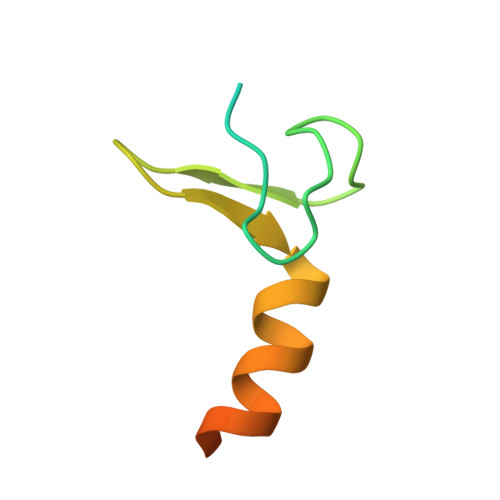
Solution structure of the dimeric zinc binding domain of the chaperone ClpX.
Donaldson, L.W., Wojtyra, U., Houry, W.A.(2003) J Biological Chem 278: 48991-48996
- PubMed: 14525985 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M307826200
- Primary Citation Related Structures:
1OVX - PubMed Abstract:
ClpX (423 amino acids), a member of the Clp/Hsp100 family of molecular chaperones and the protease, ClpP, comprise a multimeric complex supporting targeted protein degradation in Escherichia coli. The ClpX sequence consists of an NH2-terminal zinc binding domain (ZBD) and a COOH-terminal ATPase domain. Earlier, we have demonstrated that the zinc binding domain forms a constitutive dimer that is essential for the degradation of some ClpX substrates such as gammaO and MuA but is not required for the degradation of other substrates such as green fluorescent protein-SsrA. In this report, we present the NMR solution structure of the zinc binding domain dimer. The monomer fold reveals that ZBD is a member of the treble clef zinc finger family, a motif known to facilitate protein-ligand, protein-DNA, and protein-protein interactions. However, the dimeric ZBD structure is not related to any protein structure in the Protein Data Bank. A trimer-of-dimers model of ZBD is presented, which might reflect the closed state of the ClpX hexamer.
- Department of Biology, York University, Toronto, Ontario M3J 1P3, Canada. logand@yorku.ca
Organizational Affiliation: